| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 10,015,575 – 10,015,681 |
| Length | 106 |
| Max. P | 0.950160 |

| Location | 10,015,575 – 10,015,681 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 82.95 |
| Mean single sequence MFE | -49.23 |
| Consensus MFE | -34.09 |
| Energy contribution | -37.40 |
| Covariance contribution | 3.31 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.950160 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 10015575 106 - 23771897 GUCAAGGCGGUCAGCUGAU-GGGGAAUUGC-AGAGUCGGAGGUCACCAGUCGGCGGUGGUCACUGUCCACUGGUGUUCCUCUGGACUUGGCUGC--AGCCCCCGGGCCAA .....(((.(((....)))-((((..((((-(((((((((((.(((((((.((((((....)))))).)))))))..))))).))))...))))--)).))))..))).. ( -52.60) >DroSec_CAF1 68434 92 - 1 GUCAAGGCGGUCAGCUGGU-GGGGAAUAGC-GGGGUCGGAGGUCACCAGUCG---GUGGUCACUGUCCAAUGGUGGACCUCUGGAC-------------CCCCGGGCCAA ((....))((((.((((..-......))))-(((((((((((((((((..((---((....)))).....)))).))))))).)))-------------)))..)))).. ( -42.70) >DroSim_CAF1 78098 103 - 1 GUCAAGGCGGUCAGCUGGU-GGGGAGUAGC-GGGGUCGGAGGUCACCAGUCG---GUGGUCACUGUCCACUGGUGGACCUCUGGACUUGGCUGC--AGCCCCCGGGCCAA ((....))(((...((((.-(((..(((((-.((((((((((((((((((.(---(..(...)..)).)))))).))))))).))))).)))))--..)))))))))).. ( -53.60) >DroYak_CAF1 70090 110 - 1 GUCAAGGCGGUCAGCUGUUUGGGGAGUUGGAGGAUUCGGAGGUCACCAGUCGGCAGUGGUCACUGUACACUGGUGGACCUAUGGAUUUGGAACCCGAGGCCCCGGGCCAA .....(((.....)))((((((((..((((.(((((((.(((((((((((..(((((....)))))..)))))).))))).))))))).....))))..))))))))... ( -48.00) >consensus GUCAAGGCGGUCAGCUGGU_GGGGAAUAGC_GGAGUCGGAGGUCACCAGUCG___GUGGUCACUGUCCACUGGUGGACCUCUGGACUUGGCUGC__AGCCCCCGGGCCAA .....(((((....))....((((....((..((((((((((((((((((.((((((....)))))).))))))).)))))).)))))....)).....))))..))).. (-34.09 = -37.40 + 3.31)
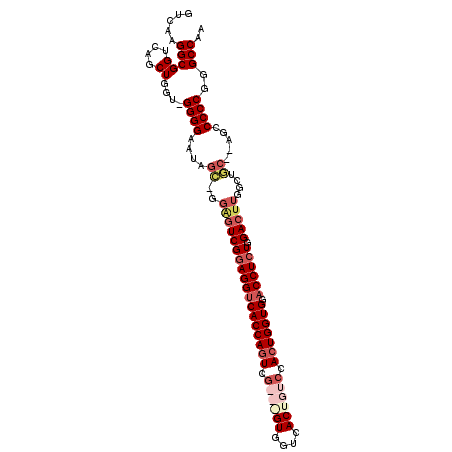



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:57:36 2006