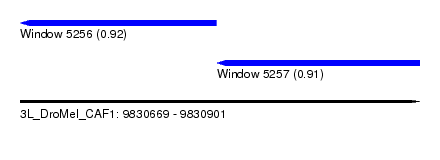
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,830,669 – 9,830,901 |
| Length | 232 |
| Max. P | 0.918934 |

| Location | 9,830,669 – 9,830,783 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.56 |
| Mean single sequence MFE | -38.17 |
| Consensus MFE | -32.26 |
| Energy contribution | -33.01 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.918934 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
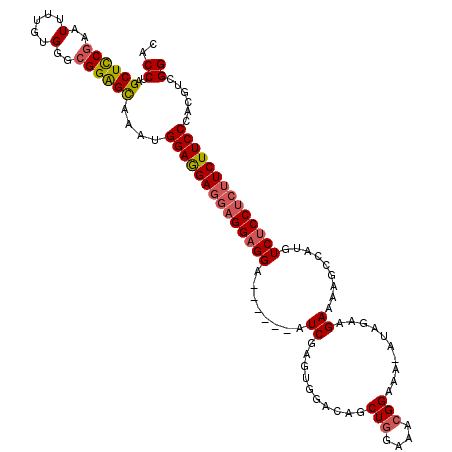
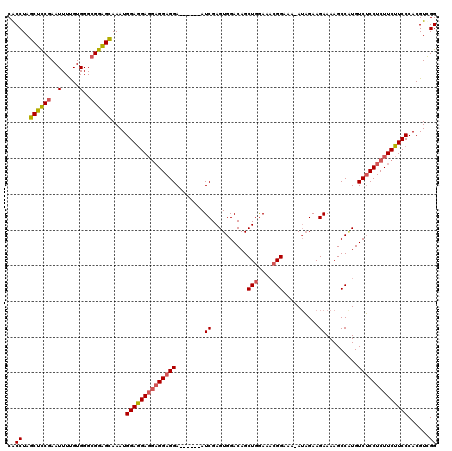
>3L_DroMel_CAF1 9830669 114 - 23771897 CGCCUAGCUUCGAAUUUUGUGGGCGGGGCAAAUGGAGGAGGAGGAGGA------AUCGAGUGGACAGCUGGAAACGGAAAAAAAGAAGAAAAGCCCUGUCUCCUCUUCUUCCCACGUCGG .((((.(((((((...))).)))).))))..(((..((((((((((((------..(.((.((....(((....)))................)))))..))))))))))))..)))... ( -38.45) >DroSec_CAF1 124638 110 - 1 CACCUAGCUCCGAAUUUUGUGGGCGGAGCAAAUGGAGGA---GGAGGA------AUCGAGUGGACAGCUGGAAACGGAAA-AUAGAAGAAAAGCCAUGUCUCCUCUUCUUCCCACGUCGG ..((..((((((..(.....)..))))))....((((((---((((((------.....((((....(((....)))...-............))))...))))))))))))......)) ( -38.91) >DroSim_CAF1 127950 113 - 1 CACCUAGCUCCGAAUUUUGUGGGCGGAGCAAAUGGAGGAGGAGGAGGA------AUCGAGUGGACAGCUGGAAACGGAAA-AUAGAAGAAAAGCCAUGUCUCCUCUUCUUCCCACGUCGG ..((..((((((..(.....)..))))))....((((((((((((((.------.....((((....(((....)))...-............)))).))))))))))))))......)) ( -42.21) >DroYak_CAF1 137891 119 - 1 CACCUAGCUCCAAAUUUCGUGGGAGGAGUAAAUGGAAGAGGAGGAGGAGGGAGGAUCGAGUGGACAGCUGAAAUAGGAAA-AUUGAAGAAAAGCCGUGUCGCCUCUUCUUCCCACGUUGG ..(((..((((...((((((...........))))))..)))).))).(((((((..(((..(((((((...........-..........)))..))))..))).)))))))....... ( -33.10) >consensus CACCUAGCUCCGAAUUUUGUGGGCGGAGCAAAUGGAGGAGGAGGAGGA______AUCGAGUGGACAGCUGGAAACGGAAA_AUAGAAGAAAAGCCAUGUCUCCUCUUCUUCCCACGUCGG ..((..((((((..(.....)..))))))....((((((((((((((........((..........(((....)))..........)).........))))))))))))))......)) (-32.26 = -33.01 + 0.75)

| Location | 9,830,783 – 9,830,901 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.28 |
| Mean single sequence MFE | -25.55 |
| Consensus MFE | -21.55 |
| Energy contribution | -22.05 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.910318 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
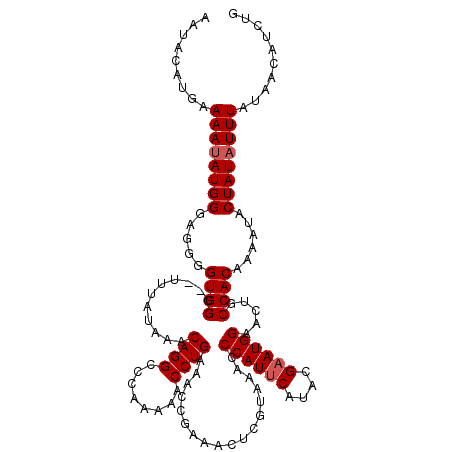
>3L_DroMel_CAF1 9830783 118 - 23771897 AAUAUGAGAAAAUAUGGGAGGAGUGGG--UUUAUAAACAGGCCCAAAAACCUGAAACCGAAACUCGUAGACCCAUUCAUACGAAUGGAAAUGCCACAAAAUACUAUAUUUAUAACAUCUG .........((((((((...(((((((--(((((...((((........)))).....(.....))))))))))))).......(((.....))).......)))))))).......... ( -28.10) >DroSec_CAF1 124748 118 - 1 AAUACAUGAAAAUAUGGGAGGGGUGGU--CUUAUAAACAGGCCCAAAAACCUGAAACCGAAACUCGUAAACCCAUUCAUACGAAUGGAUCUGCCACAAAAUACUAUAUUUAUAACAUCUG .........((((((((.....(((((--........((((........))))..................((((((....))))))....)))))......)))))))).......... ( -25.00) >DroSim_CAF1 128063 118 - 1 AAUACAUGAAAAUAUGGGAGGGGUGGG--UUCAUAAACAGGCCCAAAAACCUGAAACCGAAACUCGUAAACCCAUUCAUACGAAUGGAACUGCCACAAAAUACUAUAUUUAUAACAUCUG .........((((((((.(((..((((--((........))))))....))).............(((...((((((....))))))...))).........)))))))).......... ( -24.50) >DroYak_CAF1 138010 120 - 1 AAUACAUGAAAAUAUGGAAGGGGUGGGGUUUUAUAAACAGGCCCGAAAACCUGAAACCGAAACUCGUAAACCCAUUCAUACGCAUGGAACUGCCACAAAAUACUAUUUUUAUAACAUCUG .....(((((((((..(((.((((.(((((((.....((((........)))).....)))))))....)))).)))....(((......)))..........)))))))))........ ( -24.60) >consensus AAUACAUGAAAAUAUGGGAGGGGUGGG__UUUAUAAACAGGCCCAAAAACCUGAAACCGAAACUCGUAAACCCAUUCAUACGAAUGGAACUGCCACAAAAUACUAUAUUUAUAACAUCUG .........((((((((.....((((...........((((........))))..................((((((....)))))).....))))......)))))))).......... (-21.55 = -22.05 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:56:17 2006