
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,827,395 – 9,827,551 |
| Length | 156 |
| Max. P | 0.987926 |

| Location | 9,827,395 – 9,827,511 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.64 |
| Mean single sequence MFE | -30.52 |
| Consensus MFE | -27.09 |
| Energy contribution | -27.27 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.13 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.10 |
| SVM RNA-class probability | 0.987926 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9827395 116 + 23771897 UGGGAAACGGAAAUCAAAAGCGGCAAGGCAACAAUGUUGUUGCAGCUUUGCAUAUAAUUUA----UGUUGAAAAAGCAUAGCAAAAGCUGCUUAUGAGUUGCGAGAUUGAAAAUGUGAGA .......((.((.(((.(((((((...((((((....)))))).((((((((((.....))----)))....))))).........))))))).))).)).))................. ( -31.80) >DroSec_CAF1 121365 116 + 1 UGGGAAACGGAAAUCAAAAGCGGCAAGGCAACAAUGUUGUUGCAGCUUUGCAUAUAAUUUA----UGUUGAAAAAGUAUAGCAAAAGCUGCUUAUGAGUUGCGAGAUUGAAAAUGUGAGA .......((.((.(((.(((((((...((((((....)))))).((((((((((.....))----)))....))))).........))))))).))).)).))................. ( -29.50) >DroSim_CAF1 124652 116 + 1 UGGGAAACGGAAAUCAAAAGCGGCAAGGCAGCAAUGUUGUUGCAGCUUUGCAUAUAAUUUA----UGUUGAAAAAGCAUAGCAAAAGCUGCUUAUGAGUUGCGAGAUUGAAAAUGUGAGA .......((.((.(((.(((((((...((((((....)))))).((((((((((.....))----)))....))))).........))))))).))).)).))................. ( -31.80) >DroYak_CAF1 134500 115 + 1 UGGGAAACGGAAAUCAAAAGCGGC-----AACAAUGUUGUUGCAGCUUUGCAUAUAAUUUAUGUAUGUUGAAGAAGCAUAGCAAAAUCCGCUUAUGAGUUGCGAGAUUGAAAAUGUGAGG .......((.((.(((.(((((((-----((((....)))))).((((((((((((.....)))))))....)))))...........))))).))).)).))................. ( -29.00) >consensus UGGGAAACGGAAAUCAAAAGCGGCAAGGCAACAAUGUUGUUGCAGCUUUGCAUAUAAUUUA____UGUUGAAAAAGCAUAGCAAAAGCUGCUUAUGAGUUGCGAGAUUGAAAAUGUGAGA .......((.((.(((.(((((((...((((((....)))))).(((.(((...((((........)))).....))).)))....))))))).))).)).))................. (-27.09 = -27.27 + 0.19)


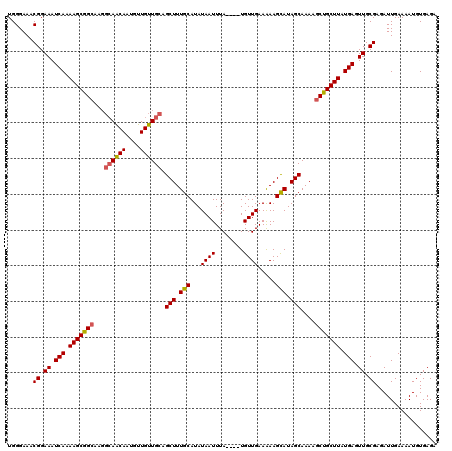
| Location | 9,827,395 – 9,827,511 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.64 |
| Mean single sequence MFE | -21.89 |
| Consensus MFE | -16.62 |
| Energy contribution | -17.50 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.04 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.759635 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9827395 116 - 23771897 UCUCACAUUUUCAAUCUCGCAACUCAUAAGCAGCUUUUGCUAUGCUUUUUCAACA----UAAAUUAUAUGCAAAGCUGCAACAACAUUGUUGCCUUGCCGCUUUUGAUUUCCGUUUCCCA .................((.((.(((.((((((((((.((((((.(((.......----.))).)))).))))))))((((((....))))))......)))).))).)).))....... ( -22.80) >DroSec_CAF1 121365 116 - 1 UCUCACAUUUUCAAUCUCGCAACUCAUAAGCAGCUUUUGCUAUACUUUUUCAACA----UAAAUUAUAUGCAAAGCUGCAACAACAUUGUUGCCUUGCCGCUUUUGAUUUCCGUUUCCCA .................((.((.(((.((((((((((.((((((.(((.......----.))).)))).))))))))((((((....))))))......)))).))).)).))....... ( -23.10) >DroSim_CAF1 124652 116 - 1 UCUCACAUUUUCAAUCUCGCAACUCAUAAGCAGCUUUUGCUAUGCUUUUUCAACA----UAAAUUAUAUGCAAAGCUGCAACAACAUUGCUGCCUUGCCGCUUUUGAUUUCCGUUUCCCA .................((.((.(((.(((((((....)))..............----..........((((.((.((((.....)))).)).)))).)))).))).)).))....... ( -20.40) >DroYak_CAF1 134500 115 - 1 CCUCACAUUUUCAAUCUCGCAACUCAUAAGCGGAUUUUGCUAUGCUUCUUCAACAUACAUAAAUUAUAUGCAAAGCUGCAACAACAUUGUU-----GCCGCUUUUGAUUUCCGUUUCCCA .................((.((.(((.(((((......(((.(((........................))).))).((((((....))))-----))))))).))).)).))....... ( -21.26) >consensus UCUCACAUUUUCAAUCUCGCAACUCAUAAGCAGCUUUUGCUAUGCUUUUUCAACA____UAAAUUAUAUGCAAAGCUGCAACAACAUUGUUGCCUUGCCGCUUUUGAUUUCCGUUUCCCA .................((.((.(((.((((((((((.((((((.(((............))).)))).))))))))((((((....))))))......)))).))).)).))....... (-16.62 = -17.50 + 0.88)



| Location | 9,827,435 – 9,827,551 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.94 |
| Mean single sequence MFE | -16.17 |
| Consensus MFE | -13.23 |
| Energy contribution | -13.68 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.506220 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9827435 116 - 23771897 UUUCACACCUGCACACGCAUUUAUUUUUACCUCUUCUGCUUCUCACAUUUUCAAUCUCGCAACUCAUAAGCAGCUUUUGCUAUGCUUUUUCAACA----UAAAUUAUAUGCAAAGCUGCA .........(((....))).................(((...................)))........((((((((.((((((.(((.......----.))).)))).)))))))))). ( -17.21) >DroSim_CAF1 124692 116 - 1 UUUCACACCUGCACACGCAUUUAUUUUUACCUCUUCUGCUUCUCACAUUUUCAAUCUCGCAACUCAUAAGCAGCUUUUGCUAUGCUUUUUCAACA----UAAAUUAUAUGCAAAGCUGCA .........(((....))).................(((...................)))........((((((((.((((((.(((.......----.))).)))).)))))))))). ( -17.21) >DroYak_CAF1 134535 120 - 1 UUUCACACCUGCACACGCAUUUAUUUUUACCUCUUCUGCGCCUCACAUUUUCAAUCUCGCAACUCAUAAGCGGAUUUUGCUAUGCUUCUUCAACAUACAUAAAUUAUAUGCAAAGCUGCA ..........(((..((((.................))))............((((.(((.........)))))))..(((.(((........................))).)))))). ( -14.09) >consensus UUUCACACCUGCACACGCAUUUAUUUUUACCUCUUCUGCUUCUCACAUUUUCAAUCUCGCAACUCAUAAGCAGCUUUUGCUAUGCUUUUUCAACA____UAAAUUAUAUGCAAAGCUGCA .........(((....))).................(((...................)))........((((((((.((((((.(((............))).)))).)))))))))). (-13.23 = -13.68 + 0.45)

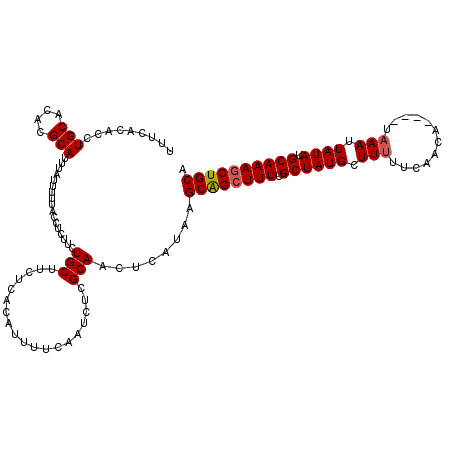

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:56:11 2006