
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,825,027 – 9,825,251 |
| Length | 224 |
| Max. P | 0.994968 |

| Location | 9,825,027 – 9,825,124 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 97.13 |
| Mean single sequence MFE | -35.18 |
| Consensus MFE | -32.92 |
| Energy contribution | -32.44 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.94 |
| SVM decision value | 2.34 |
| SVM RNA-class probability | 0.992604 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
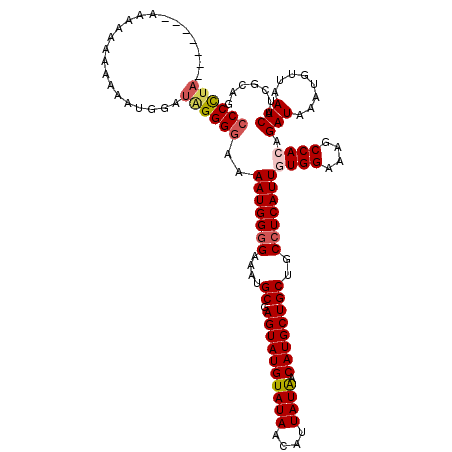
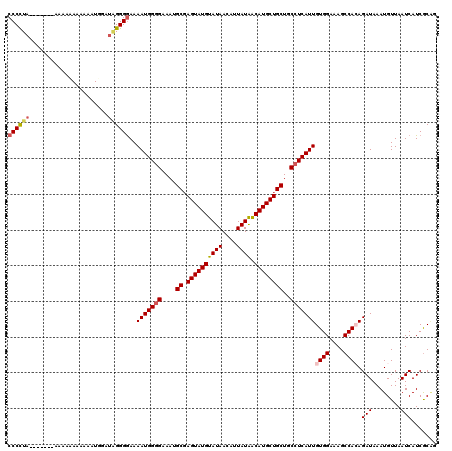
>3L_DroMel_CAF1 9825027 97 - 23771897 CUUGACCUCCUCGCAUUUACUUUGAUGUUGUUGACUUCAU-CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAGUUGUCGUCCGGAUCGAGU .((((((..((((..(((((...(((((..((((((((..-..))))))))..))))).)))))....))))..))))))...(((.......))).. ( -34.20) >DroSec_CAF1 119051 97 - 1 CUUGACCUCUUCGCAUUUACUUUGAUGUUGUUGACUUCAU-CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAGUUGUCGUCCGGAUCGGAU .(((((((.((((..(((((...(((((..((((((((..-..))))))))..))))).)))))....)))).))))))).....(((((...))))) ( -34.10) >DroSim_CAF1 122313 97 - 1 CUUGACCUCCUCGCAUUUACUUUGAUGUUGUUGACUUCAU-CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAGUUGUCGUCCGGAUCGGAU .((((((..((((..(((((...(((((..((((((((..-..))))))))..))))).)))))....))))..)))))).....(((((...))))) ( -36.90) >DroEre_CAF1 121885 97 - 1 CUUGACCUCCUCGCAUUUACUUUGAUGUUGUUGACUUCAU-CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAGUUGUCGCGCGGAUCGGAU .((((((..((((..(((((...(((((..((((((((..-..))))))))..))))).)))))....))))..))))))..(((.((.....))))) ( -33.60) >DroYak_CAF1 132094 98 - 1 CUUGACCUCCUCGCAUUUGCUUUGAUGUUGUUGACUUCAUUCAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAGUUGUCGUCCGGAUCGGAU .((((((..((((..((..(...(((((..((((((((.....))))))))..))))).)..))....))))..)))))).....(((((...))))) ( -37.10) >consensus CUUGACCUCCUCGCAUUUACUUUGAUGUUGUUGACUUCAU_CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAGUUGUCGUCCGGAUCGGAU .((((((..((((..(((((...(((((..((((((((.....))))))))..))))).)))))....))))..))))))...(((.......))).. (-32.92 = -32.44 + -0.48)



| Location | 9,825,045 – 9,825,151 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 91.44 |
| Mean single sequence MFE | -34.08 |
| Consensus MFE | -32.94 |
| Energy contribution | -32.75 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.594077 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9825045 106 - 23771897 GGGGGGG---GCU---UUUUUUAUGACUUAACGCUUGACCUCCUCGCAUUUACUUUGAUGUUGUUGACUUCAU-CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAG (.(..((---((.---........).)))..).).(((((..((((..(((((...(((((..((((((((..-..))))))))..))))).)))))....))))..))))). ( -34.60) >DroSim_CAF1 122331 109 - 1 GGAUGGGGAGCCC---UUUUUUAUGACUUAACGCUUGACCUCCUCGCAUUUACUUUGAUGUUGUUGACUUCAU-CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAG ....(((....))---)..................(((((..((((..(((((...(((((..((((((((..-..))))))))..))))).)))))....))))..))))). ( -35.50) >DroEre_CAF1 121903 101 - 1 G--------GCCC---UUUUUUAUGACUUAACGCUUGACCUCCUCGCAUUUACUUUGAUGUUGUUGACUUCAU-CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAG .--------....---...................(((((..((((..(((((...(((((..((((((((..-..))))))))..))))).)))))....))))..))))). ( -33.00) >DroYak_CAF1 132112 105 - 1 G--------GCCCCUUUUUUUUAUGACUUAACGCUUGACCUCCUCGCAUUUGCUUUGAUGUUGUUGACUUCAUUCAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAG .--------..........................(((((..((((..((..(...(((((..((((((((.....))))))))..))))).)..))....))))..))))). ( -33.20) >consensus G________GCCC___UUUUUUAUGACUUAACGCUUGACCUCCUCGCAUUUACUUUGAUGUUGUUGACUUCAU_CAGGGGUCAGAGGCAUCUGUGGAACACCGGGUGGGUCAG ...................................(((((..((((..(((((...(((((..((((((((.....))))))))..))))).)))))....))))..))))). (-32.94 = -32.75 + -0.19)



| Location | 9,825,151 – 9,825,251 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.87 |
| Mean single sequence MFE | -28.20 |
| Consensus MFE | -24.17 |
| Energy contribution | -24.81 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.53 |
| SVM RNA-class probability | 0.994968 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9825151 100 + 23771897 CCCCUA-------AAAA-------------GGGAAAAAUGGGGAAAUGCGAGUAUGUAUAACAUUAUAACAUGCUGCUGCCUCAUUGUGGAAAGCCACAGAUAAAUGUUAAUCAUCACAG .((((.-------...)-------------)))...(((((((....((.((((((((((....))).)))))))))..)))))))((((....)))).(((........)))....... ( -27.90) >DroSec_CAF1 119185 113 + 1 CCCCUA-------AAAAUAAAAAAUGGAUGGGGGAAAAUGGGGAAAUGCGAGUAUGUAUAACAUUAUAACAUGCUGCUGCCUCAUUGUGGAAAGCCACAGAUAAAUGUUAAUCAUCGCAG ((((((-------...((.....))...))))))..(((((((....((.((((((((((....))).)))))))))..)))))))((((....)))).(((........)))....... ( -30.70) >DroSim_CAF1 122440 113 + 1 CCCUUA-------AAAAGAAAAAAUGGAUGGGGGAAAAUGGGGAAAUGCGAGUAUGUAUAACAUUAUAACAUGCUGCUGCAUCAUUGUGGAAAGCCACAGAUAAAUGUUAAUCAUCGCAG ((((((-------...............))))))............(((((...((..(((((((.....((((....))))..((((((....))))))...)))))))..))))))). ( -25.26) >DroEre_CAF1 122004 120 + 1 CCCCUACAAAAAAAAAAAAAAAAAUGGCUAGGGGAAAAUGGGGAAAUGCGAGUAUGUAUAACAUUAUGACAUGCUGCUGCCUCAUUGUGGAAAGCCAGAGAUAAAUGUUAAUCAUCGCUG ((((((......................))))))..(((((((....((.((((((((((....))).)))))))))..))))))).(((....)))(((((........))).)).... ( -27.45) >DroYak_CAF1 132217 110 + 1 CCCCUA----------AAAAAAAAUUGGUAGGGGAAAAUGGGGAAAUGCCAGUAUGUAUAACAUUAUGGCAUGCUGCUGCCUCAUUGUGGAAAGCCAGAGAUAAAUGUUAAUCAUCGCGG ((((((----------..((....))..))))))..(((((((....((.((((((((((....))).)))))))))..))))))).(((....)))(((((........))).)).... ( -29.70) >consensus CCCCUA_______AAAAAAAAAAAUGGAUAGGGGAAAAUGGGGAAAUGCGAGUAUGUAUAACAUUAUAACAUGCUGCUGCCUCAUUGUGGAAAGCCACAGAUAAAUGUUAAUCAUCGCAG ((((((......................))))))..(((((((....((.((((((((((....)))).))))))))..)))))))((((....)))).(((........)))....... (-24.17 = -24.81 + 0.64)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:56:03 2006