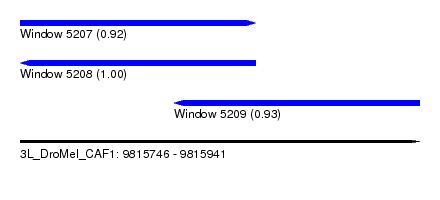

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,815,746 – 9,815,941 |
| Length | 195 |
| Max. P | 0.998829 |

| Location | 9,815,746 – 9,815,861 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.70 |
| Mean single sequence MFE | -22.68 |
| Consensus MFE | -20.22 |
| Energy contribution | -20.46 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924006 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9815746 115 + 23771897 UUGCCUUACUACC----UUCA-AAUGGCAAUAACAAUAGCGUACGCAAAGCGGAGCUCCGCCAGUUCAUCUGCAUAAUAAUGCUAAUUAUUUUUAAUUGCCGCAACUUGUCAUUUGCCAC .............----....-..((((((..((((..(((...((((.((((....))))..........((((....)))).............)))))))...))))...)))))). ( -25.60) >DroSec_CAF1 109874 115 + 1 UUAUCUUACUACC----UCCA-AAUAGCAAUAACAAUAGCGUACGCAAAGCGGAGCUCCGCCAGUUCAUCUGCAUAAUAAUGCUAAUUAUUUUUAAUUGCCGCAACUUGUCAUUUGCCAC .............----....-....((((..((((..(((...((((.((((....))))..........((((....)))).............)))))))...))))...))))... ( -20.90) >DroSim_CAF1 113208 116 + 1 UUGUCUUACUACC----UCCANAAUAGCAAUAACAAUAGCGUACGCAAAGCGGAGCUUCGCCAGUUCAUUUGCAUAAUAAUGCUAAUUAUUUUUAAUUGCCGCAACUUGUCAUUUGCCAC .............----.........((((..((((..(((...(((((...(((((.....))))).)))))........((.(((((....))))))))))...))))...))))... ( -21.10) >DroEre_CAF1 112711 115 + 1 UUGUCUUACCACC----UCCA-AAUGGCAAUAACAAUAGCGUACGCAAAGCGGAGCUUCGCCAGUUCAUCUGCAUAAUAAUACUAAUUAUUUUUAAUUGCCGCAACUUGUCAUUUGCCAC .............----....-..((((((..((((..(((...((((.(((((....(....)....)))))..((((((....)))))).....)))))))...))))...)))))). ( -22.90) >DroYak_CAF1 122788 119 + 1 UUGUCUUACUACCGCCCUCAA-AAUGGCAAUAACAAUAGCGUACGCAAAGCGGAGCUUCGCCAGUUCAUCUGCAUAAUAAUACUAAUUAUUUUUAAUUGCCGCAACUUGUCAUUUGCCAC .....................-..((((((..((((..(((...((((.(((((....(....)....)))))..((((((....)))))).....)))))))...))))...)))))). ( -22.90) >consensus UUGUCUUACUACC____UCCA_AAUGGCAAUAACAAUAGCGUACGCAAAGCGGAGCUUCGCCAGUUCAUCUGCAUAAUAAUGCUAAUUAUUUUUAAUUGCCGCAACUUGUCAUUUGCCAC ........................((((((..((((..(((...((((.(((((....(....)....)))))..((((((....)))))).....)))))))...))))...)))))). (-20.22 = -20.46 + 0.24)


| Location | 9,815,746 – 9,815,861 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.70 |
| Mean single sequence MFE | -31.48 |
| Consensus MFE | -30.84 |
| Energy contribution | -30.12 |
| Covariance contribution | -0.72 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.98 |
| SVM decision value | 3.24 |
| SVM RNA-class probability | 0.998829 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9815746 115 - 23771897 GUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGCAUUAUUAUGCAGAUGAACUGGCGGAGCUCCGCUUUGCGUACGCUAUUGUUAUUGCCAUU-UGAA----GGUAGUAAGGCAA (((((((.((((((...((((((((((....))))).)).....(((((((.......(((((....)))))))))))))))..))))))))))))).-....----............. ( -34.21) >DroSec_CAF1 109874 115 - 1 GUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGCAUUAUUAUGCAGAUGAACUGGCGGAGCUCCGCUUUGCGUACGCUAUUGUUAUUGCUAUU-UGGA----GGUAGUAAGAUAA (((((((.((((((...((((((((((....))))).)).....(((((((.......(((((....)))))))))))))))..))))))))))))).-....----............. ( -31.91) >DroSim_CAF1 113208 116 - 1 GUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGCAUUAUUAUGCAAAUGAACUGGCGAAGCUCCGCUUUGCGUACGCUAUUGUUAUUGCUAUUNUGGA----GGUAGUAAGACAA (((((((.((((((...(((((...............((((....))))..........(((((((...))))))))).)))..)))))))))))))......----............. ( -29.70) >DroEre_CAF1 112711 115 - 1 GUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGUAUUAUUAUGCAGAUGAACUGGCGAAGCUCCGCUUUGCGUACGCUAUUGUUAUUGCCAUU-UGGA----GGUGGUAAGACAA (((((((.((((((...(((((.............((((.(((((.....)))))))))(((((((...))))))))).)))..))))))))))))).-....----............. ( -30.80) >DroYak_CAF1 122788 119 - 1 GUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGUAUUAUUAUGCAGAUGAACUGGCGAAGCUCCGCUUUGCGUACGCUAUUGUUAUUGCCAUU-UUGAGGGCGGUAGUAAGACAA (((((((.((((((...(((((.............((((.(((((.....)))))))))(((((((...))))))))).)))..))))))))))))).-..................... ( -30.80) >consensus GUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGCAUUAUUAUGCAGAUGAACUGGCGAAGCUCCGCUUUGCGUACGCUAUUGUUAUUGCCAUU_UGGA____GGUAGUAAGACAA (((((((.((((((...(((((...............((((....))))..........(((((((...))))))))).)))..)))))))))))))....................... (-30.84 = -30.12 + -0.72)



| Location | 9,815,821 – 9,815,941 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.61 |
| Mean single sequence MFE | -29.67 |
| Consensus MFE | -25.14 |
| Energy contribution | -25.58 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928082 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9815821 120 - 23771897 GCCCAUCCUCGGCGACAGUGAAAACCGAACCUCCAGUCAGUUGCCCGCGAAACUUUCACUAAUCACUCACAGAGUUUUGUGUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGCA ((((......)(((((.((....))..........((((.((((((((((((((((..............))))))))))).))))).))))..)))))))).................. ( -34.34) >DroSec_CAF1 109949 119 - 1 GCCAAUCCUAGGCGACAGUGAAAACCGAACCUC-AGUCAGUUGCCCGCGAAACUUUCACUAAUCACUCACAGAGUUUUGUGUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGCA ...........(((((.((....))........-.((((.((((((((((((((((..............))))))))))).))))).))))..))))).(((((((....))))).)). ( -34.04) >DroSim_CAF1 113284 119 - 1 GCCAAUCCUCGGCGACAGUGAAAACCGAACCUC-AAUCAGUUGCCCGCGAAACUUUCACUAAUCACUCACAGAGUUUUGUGUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGCA ...........(((((.((....))........-..(((.((((((((((((((((..............))))))))))).))))).)))...))))).(((((((....))))).)). ( -30.24) >DroEre_CAF1 112786 115 - 1 GCUUAUCCUCUCCGACUGUGAAAACCGAACCU-----CAGUUGCCCGAGAAACUUUCACUAAUCACUCACAGAGUUUUGUGUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGUA ........((((((((((.(..........).-----))))))...)))).......((((((.((((...))))....(((.((((........)))).)))..........)))))). ( -20.00) >DroYak_CAF1 122867 115 - 1 GCCCAUCGUCUGCGACAGUGGAAACCGUAGCU-----CAGUUGCCCGAGAAACUUUCACUAAUCACUCACAGAGUUUUGUGUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGUA ..((((.(((...))).))))...((((((((-----((.(((((((.((((((((..............)))))))).)).))))).)))...)))))))................... ( -29.74) >consensus GCCAAUCCUCGGCGACAGUGAAAACCGAACCUC_A_UCAGUUGCCCGCGAAACUUUCACUAAUCACUCACAGAGUUUUGUGUGGCAAAUGACAAGUUGCGGCAAUUAAAAAUAAUUAGCA (((........(((((.((....))............((.((((((((((((((((..............))))))))))).))))).))....)))))))).................. (-25.14 = -25.58 + 0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:55:31 2006