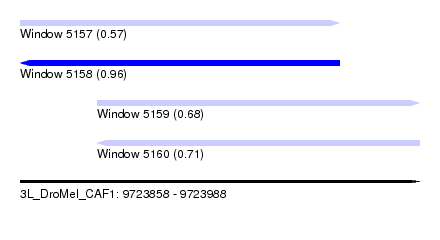
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,723,858 – 9,723,988 |
| Length | 130 |
| Max. P | 0.964485 |

| Location | 9,723,858 – 9,723,962 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 74.37 |
| Mean single sequence MFE | -21.34 |
| Consensus MFE | -11.14 |
| Energy contribution | -12.06 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.573373 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9723858 104 + 23771897 CUUCUACACUUCCCCCAAUUAGAGAAA-AUAAACCAAAUGCUCUUUUAUGAAAUAAGGGUGGCUAAGUCACAAUGGUACUUAACCACCCGGUUAGUAAAAUUUGC .......(((..((.((...((((...-((.......)).))))....))......((((((.(((((.((....))))))).))))))))..)))......... ( -22.80) >DroSec_CAF1 19122 104 + 1 CUUCUACACUUUCCCCAAUUAGAGAAA-AUAAACCAAAUGCUCUUUUAUAAAAUAAGGGUGGCUAAGUCACAAUGGUAUUUAACCACCCGGUUAGUAAUAUGUGC .......(((..((......((((...-((.......)).))))............((((((.(((((.((....))))))).))))))))..)))......... ( -19.90) >DroSim_CAF1 19632 104 + 1 CUUCUACACUUUCCCCAAUUAGAGAAA-AUAAACUAAAUGCUCUUUUAUGAAAUAAGGGUGGCUAAGUCACACUGGUAUUUAACCACCCGGUUAGUAAAAUUUGC .......(((..((.((...((((...-((.......)).))))....))......((((((.(((((.((....))))))).))))))))..)))......... ( -20.20) >DroEre_CAF1 18372 98 + 1 CCUUUA------CCCCAGUUGGAGUAAACUAAACCAAAUGCUC-UUUAUGAAAUAAGGGUGGCUUAGUGAUAUUGGCACUUAAGCACCUUGUUAGUAAGCCGUGC ((((((------...((...((((((............)))))-)...))...)))))).((((((.(((((..(((......)).)..))))).)))))).... ( -26.10) >DroYak_CAF1 27758 79 + 1 --UUUC------UACCAAUUAAAG------AAAGCAAAAGCUGUUUUAUGAAAUAAGGGUGGUGU------------UUUCAACCACCUGGUUAGUGAACCUUGC --.(((------.(((.....(((------(.(((....))).)))).........(((((((..------------.....))))))))))....)))...... ( -17.70) >consensus CUUCUACACUU_CCCCAAUUAGAGAAA_AUAAACCAAAUGCUCUUUUAUGAAAUAAGGGUGGCUAAGUCACAAUGGUAUUUAACCACCCGGUUAGUAAAAUUUGC .......................................((...(((((.((....((((((.(((((.........))))).))))))..)).)))))....)) (-11.14 = -12.06 + 0.92)

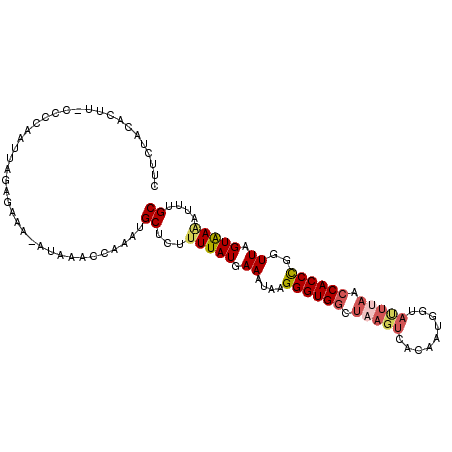
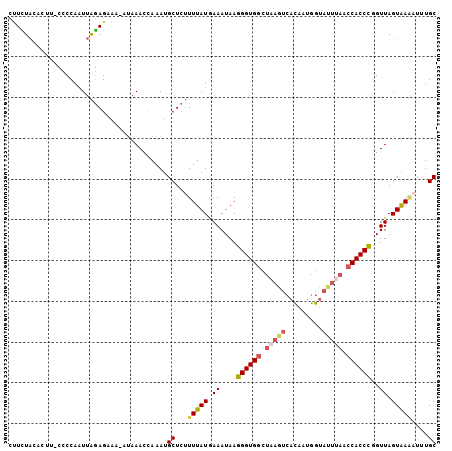
| Location | 9,723,858 – 9,723,962 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 74.37 |
| Mean single sequence MFE | -25.34 |
| Consensus MFE | -11.17 |
| Energy contribution | -13.12 |
| Covariance contribution | 1.95 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.44 |
| SVM decision value | 1.57 |
| SVM RNA-class probability | 0.964485 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9723858 104 - 23771897 GCAAAUUUUACUAACCGGGUGGUUAAGUACCAUUGUGACUUAGCCACCCUUAUUUCAUAAAAGAGCAUUUGGUUUAU-UUUCUCUAAUUGGGGGAAGUGUAGAAG .....((((((.((((((((((((((((((....)).))))))))))))....(((......))).....)))).((-(((((((....))))))))))))))). ( -33.00) >DroSec_CAF1 19122 104 - 1 GCACAUAUUACUAACCGGGUGGUUAAAUACCAUUGUGACUUAGCCACCCUUAUUUUAUAAAAGAGCAUUUGGUUUAU-UUUCUCUAAUUGGGGAAAGUGUAGAAG ((((........((((((((((((((.(((....)))..))))))))))...((((....))))......))))...-(((((((....)))))))))))..... ( -26.90) >DroSim_CAF1 19632 104 - 1 GCAAAUUUUACUAACCGGGUGGUUAAAUACCAGUGUGACUUAGCCACCCUUAUUUCAUAAAAGAGCAUUUAGUUUAU-UUUCUCUAAUUGGGGAAAGUGUAGAAG .....((((((.....((((((((((.(((....)))..)))))))))).............((((.....))))((-(((((((....))))))))))))))). ( -26.80) >DroEre_CAF1 18372 98 - 1 GCACGGCUUACUAACAAGGUGCUUAAGUGCCAAUAUCACUAAGCCACCCUUAUUUCAUAAA-GAGCAUUUGGUUUAGUUUACUCCAACUGGGG------UAAAGG (((((((...((....))..)))...)))).......(((((((((..(((.((....)).-)))....)))))))))(((((((....))))------)))... ( -23.80) >DroYak_CAF1 27758 79 - 1 GCAAGGUUCACUAACCAGGUGGUUGAAA------------ACACCACCCUUAUUUCAUAAAACAGCUUUUGCUUU------CUUUAAUUGGUA------GAAA-- ((..((((....)))).(((((((....------------).))))))................))(((((((..------........))))------))).-- ( -16.20) >consensus GCAAAUUUUACUAACCGGGUGGUUAAAUACCAUUGUGACUUAGCCACCCUUAUUUCAUAAAAGAGCAUUUGGUUUAU_UUUCUCUAAUUGGGG_AAGUGUAGAAG ................((((((((((.............)))))))))).......................(((((..((((((....))))))...))))).. (-11.17 = -13.12 + 1.95)


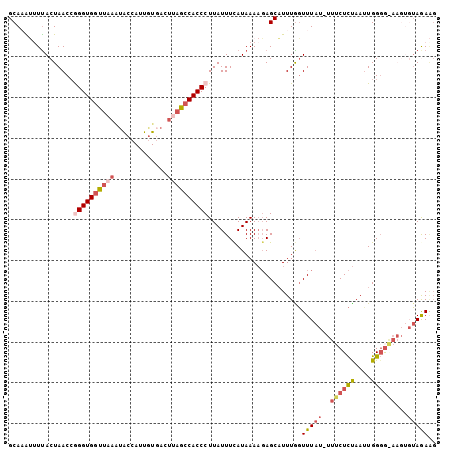
| Location | 9,723,883 – 9,723,988 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 80.83 |
| Mean single sequence MFE | -22.66 |
| Consensus MFE | -12.96 |
| Energy contribution | -14.08 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.683089 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9723883 105 + 23771897 AA-AUAAACCAAAUGCUCUUUUAUGAAAUAAGGGUGGCUAAGUCACAAUGGUACUUAACCACCCGGUUAGUAAAAUUUGCAAUUAUUAUUACCAAGAAAACCGUGC ..-..........(((..((((((.......((((((.(((((.((....))))))).)))))).....))))))...)))......................... ( -22.10) >DroSec_CAF1 19147 105 + 1 AA-AUAAACCAAAUGCUCUUUUAUAAAAUAAGGGUGGCUAAGUCACAAUGGUAUUUAACCACCCGGUUAGUAAUAUGUGCAAUUAUUAUUACCAAGAAAAUGGUGC ..-....((((.....((((...........((((((.(((((.((....))))))).))))))(((.(((((((........))))))))))))))...)))).. ( -25.90) >DroSim_CAF1 19657 105 + 1 AA-AUAAACUAAAUGCUCUUUUAUGAAAUAAGGGUGGCUAAGUCACACUGGUAUUUAACCACCCGGUUAGUAAAAUUUGCAAUUAUUAUUACCAAGAAAAUGGUGC ..-......(((.(((..((((((.......((((((.(((((.((....))))))).)))))).....))))))...))).)))....(((((......))))). ( -23.60) >DroEre_CAF1 18391 105 + 1 AAACUAAACCAAAUGCUC-UUUAUGAAAUAAGGGUGGCUUAGUGAUAUUGGCACUUAAGCACCUUGUUAGUAAGCCGUGCAAUUAUUAUUACCAAGAAUACUGUGC ..............((((-((........))))))((((((.(((((..(((......)).)..))))).))))))(..((..((((.........)))).))..) ( -21.60) >DroYak_CAF1 27774 89 + 1 -----AAAGCAAAAGCUGUUUUAUGAAAUAAGGGUGGUGU------------UUUCAACCACCUGGUUAGUGAACCUUGCAAUUAUUAUUACCAAGAAAACUGCGC -----.........((.(((((..(.((((((((((((..------------.....)))))))((((....)))).........))))).)....))))).)).. ( -20.10) >consensus AA_AUAAACCAAAUGCUCUUUUAUGAAAUAAGGGUGGCUAAGUCACAAUGGUAUUUAACCACCCGGUUAGUAAAAUUUGCAAUUAUUAUUACCAAGAAAACGGUGC .............(((...(((((.((....((((((.(((((.........))))).))))))..)).)))))....)))......................... (-12.96 = -14.08 + 1.12)


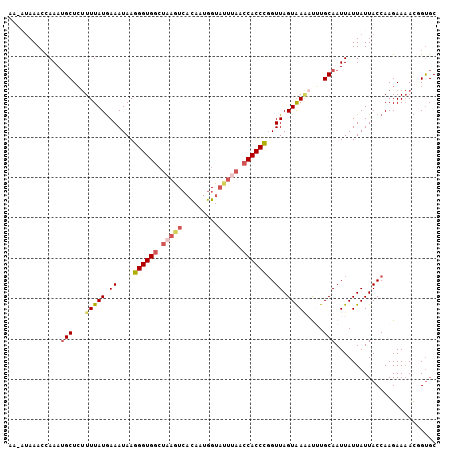
| Location | 9,723,883 – 9,723,988 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 80.83 |
| Mean single sequence MFE | -22.80 |
| Consensus MFE | -13.72 |
| Energy contribution | -14.96 |
| Covariance contribution | 1.24 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.705073 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9723883 105 - 23771897 GCACGGUUUUCUUGGUAAUAAUAAUUGCAAAUUUUACUAACCGGGUGGUUAAGUACCAUUGUGACUUAGCCACCCUUAUUUCAUAAAAGAGCAUUUGGUUUAU-UU ((..((((......(((((....)))))..........))))((((((((((((((....)).))))))))))))...............))...........-.. ( -27.49) >DroSec_CAF1 19147 105 - 1 GCACCAUUUUCUUGGUAAUAAUAAUUGCACAUAUUACUAACCGGGUGGUUAAAUACCAUUGUGACUUAGCCACCCUUAUUUUAUAAAAGAGCAUUUGGUUUAU-UU ..((((..((((((((((((...........)))))))....((((((((((.(((....)))..))))))))))...........)))))....))))....-.. ( -24.40) >DroSim_CAF1 19657 105 - 1 GCACCAUUUUCUUGGUAAUAAUAAUUGCAAAUUUUACUAACCGGGUGGUUAAAUACCAGUGUGACUUAGCCACCCUUAUUUCAUAAAAGAGCAUUUAGUUUAU-UU ..((((......)))).....(((.(((..............((((((((((.(((....)))..)))))))))).....((......))))).)))......-.. ( -22.90) >DroEre_CAF1 18391 105 - 1 GCACAGUAUUCUUGGUAAUAAUAAUUGCACGGCUUACUAACAAGGUGCUUAAGUGCCAAUAUCACUAAGCCACCCUUAUUUCAUAAA-GAGCAUUUGGUUUAGUUU (((((((((.(((((((((....)))))..(.....)...)))))))))...)))).......(((((((((..(((.((....)).-)))....))))))))).. ( -21.20) >DroYak_CAF1 27774 89 - 1 GCGCAGUUUUCUUGGUAAUAAUAAUUGCAAGGUUCACUAACCAGGUGGUUGAAA------------ACACCACCCUUAUUUCAUAAAACAGCUUUUGCUUU----- ..((.(((((..(((.(((((.........((((....)))).(((((((....------------).)))))).)))))))).))))).)).........----- ( -18.00) >consensus GCACAGUUUUCUUGGUAAUAAUAAUUGCAAAUUUUACUAACCGGGUGGUUAAAUACCAUUGUGACUUAGCCACCCUUAUUUCAUAAAAGAGCAUUUGGUUUAU_UU ((............(((((....)))))..............((((((((((.............))))))))))...............)).............. (-13.72 = -14.96 + 1.24)

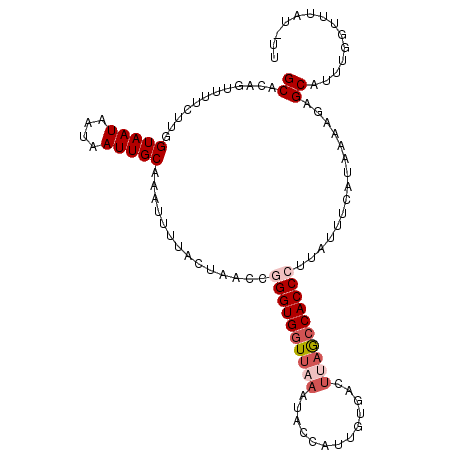

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:54:43 2006