
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,663,385 – 9,663,486 |
| Length | 101 |
| Max. P | 0.635265 |

| Location | 9,663,385 – 9,663,486 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 89.79 |
| Mean single sequence MFE | -24.72 |
| Consensus MFE | -19.71 |
| Energy contribution | -19.91 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.635265 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9663385 101 + 23771897 AAUUUCCACACUGCAAGAAAACAGAGAGCGGGGGUCAGUUGAUAGGCUGGAUGUGGGCUGGUGGUGCACUGUGGGAAAAUGUCAGCAAGA-AAACGGAUAGA ..((((((((.((((......(((...(((....((((((....)))))).)))...)))....)))).))))))))..((((.(.....-...).)))).. ( -27.40) >DroSec_CAF1 58917 99 + 1 AAUUUCCACACUGCAAGAAAACAGAGAGCGGGGGUCAGUUGAUAGGCUGCAUGUGGG--GAUGGUGCACUGUGGGAAAAUGUCAGCAAGC-AAAUGGAUAGA ..((((((((.((((......((.(..((((..(((....)))...)))).).))..--.....)))).))))))))..((((.......-.....)))).. ( -23.92) >DroSim_CAF1 61971 100 + 1 AAUUUCCACACUGCAAGAAAACAGAGAGCGGGGGUCAGUUGAUAGGCUGGAUGUGGG--GAUAGUGCACUGUGGGAAAAUGUCAGCAAGCCAAAUGGAUGGA ..((((((((.((((......(.....(((....((((((....)))))).))).).--.....)))).))))))))............(((......))). ( -23.10) >DroEre_CAF1 57865 94 + 1 AAUUUCCACACUGCAAGAAAACAGAGAGCGGGGGUCAGUUGGUAGGCUGGAU------GUGGGGUGCACUGUGGGAAAAUAUCAGCAAG--AAAUUGAUAGA ..((((((((.((((.....(((...........((((((....)))))).)------))....)))).))))))))..((((((....--...)))))).. ( -23.40) >DroYak_CAF1 59136 100 + 1 AAUUUCCACACUGCAAGAAAACAGAGAGCGGGGGUCAGUUGGUAGGCUGGAUGUGGAUGUGGGGUGCACUGUGGGAAAAUAUCAGCAAG--AAAUCGAUAGA ..((((((((.((((.....(((....(((....((((((....)))))).)))...)))....)))).))))))))..((((......--.....)))).. ( -25.80) >consensus AAUUUCCACACUGCAAGAAAACAGAGAGCGGGGGUCAGUUGAUAGGCUGGAUGUGGG__GGUGGUGCACUGUGGGAAAAUGUCAGCAAG__AAAUGGAUAGA ..((((((((.((((............(((....((((((....)))))).)))..........)))).))))))))..((((.............)))).. (-19.71 = -19.91 + 0.20)
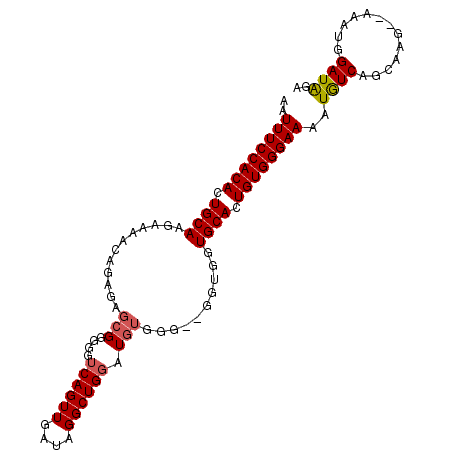



| Location | 9,663,385 – 9,663,486 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 89.79 |
| Mean single sequence MFE | -15.62 |
| Consensus MFE | -14.58 |
| Energy contribution | -14.58 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.93 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.527211 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9663385 101 - 23771897 UCUAUCCGUUU-UCUUGCUGACAUUUUCCCACAGUGCACCACCAGCCCACAUCCAGCCUAUCAACUGACCCCCGCUCUCUGUUUUCUUGCAGUGUGGAAAUU .......(((.-.......)))......(((((.((((...............(((........)))......((.....)).....)))).)))))..... ( -15.30) >DroSec_CAF1 58917 99 - 1 UCUAUCCAUUU-GCUUGCUGACAUUUUCCCACAGUGCACCAUC--CCCACAUGCAGCCUAUCAACUGACCCCCGCUCUCUGUUUUCUUGCAGUGUGGAAAUU ......((.((-(...((((.(((.........(((.......--..))))))))))....))).))....((((...((((......)))).))))..... ( -15.20) >DroSim_CAF1 61971 100 - 1 UCCAUCCAUUUGGCUUGCUGACAUUUUCCCACAGUGCACUAUC--CCCACAUCCAGCCUAUCAACUGACCCCCGCUCUCUGUUUUCUUGCAGUGUGGAAAUU .(((......)))...............(((((.((((.....--........(((........)))......((.....)).....)))).)))))..... ( -16.20) >DroEre_CAF1 57865 94 - 1 UCUAUCAAUUU--CUUGCUGAUAUUUUCCCACAGUGCACCCCAC------AUCCAGCCUACCAACUGACCCCCGCUCUCUGUUUUCUUGCAGUGUGGAAAUU ..(((((....--.....))))).....(((((.((((....((------(...(((................)))...))).....)))).)))))..... ( -16.29) >DroYak_CAF1 59136 100 - 1 UCUAUCGAUUU--CUUGCUGAUAUUUUCCCACAGUGCACCCCACAUCCACAUCCAGCCUACCAACUGACCCCCGCUCUCUGUUUUCUUGCAGUGUGGAAAUU ..(((((....--.....))))).....(((((.((((...............(((........)))......((.....)).....)))).)))))..... ( -15.10) >consensus UCUAUCCAUUU__CUUGCUGACAUUUUCCCACAGUGCACCACC__CCCACAUCCAGCCUAUCAACUGACCCCCGCUCUCUGUUUUCUUGCAGUGUGGAAAUU ............................(((((.((((...............(((........)))......((.....)).....)))).)))))..... (-14.58 = -14.58 + 0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:54:11 2006