
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,601,621 – 9,601,775 |
| Length | 154 |
| Max. P | 0.941022 |

| Location | 9,601,621 – 9,601,735 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.89 |
| Mean single sequence MFE | -19.52 |
| Consensus MFE | -18.12 |
| Energy contribution | -18.38 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.31 |
| SVM RNA-class probability | 0.941022 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9601621 114 + 23771897 CACUCUAAAAAUACAAUAAU------UCCUUACAUAAAAAUUGAAAUUAUACAAUAUAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGG ....................------.............((((........))))........(((((((((.(((..((((((((.......)))))))))))......))))))).)) ( -19.60) >DroSec_CAF1 262 114 + 1 CACUCUAAAAAUACAAUAAU------UCCUUGCAUAAACAUUAAAAUUAUACAAUAUAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGG ...............(((((------(...((......))....)))))).............(((((((((.(((..((((((((.......)))))))))))......))))))).)) ( -19.70) >DroSim_CAF1 264 114 + 1 CACUCUAAAAAUACAAUAAU------UACUUACGUAAACAUUAAAAUUAUACAAUAUAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGG ...................(------(((....))))..........................(((((((((.(((..((((((((.......)))))))))))......))))))).)) ( -20.30) >DroEre_CAF1 306 119 + 1 CACUCUGAAAC-ACAAUACUCGUAUUUCCUUAAAUCAAAAUUGUAAUAAUACAAUAAAACAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAAAGUGAUUUCCAGAGCUAUGG ......((((.-((.......)).))))...........((((((....))))))........(((((((((.(((.((.((((((.......)))))))))))......))))))).)) ( -18.50) >consensus CACUCUAAAAAUACAAUAAU______UCCUUACAUAAAAAUUAAAAUUAUACAAUAUAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGG ...............................................................(((((((((.(((..((((((((.......)))))))))))......))))))).)) (-18.12 = -18.38 + 0.25)


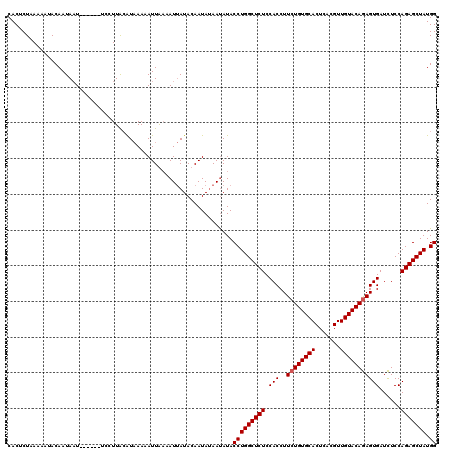
| Location | 9,601,621 – 9,601,735 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.89 |
| Mean single sequence MFE | -27.70 |
| Consensus MFE | -22.80 |
| Energy contribution | -22.55 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.506063 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9601621 114 - 23771897 CCAUAGCUCUGGAGAUCACUCUGUACAACGUGAGUGCACAGAAGGUGGAGAGCCAGGUAUAUUAUAUUGUAUAAUUUCAAUUUUUAUGUAAGGA------AUUAUUGUAUUUUUAGAGUG .....(((((((((((((((((((...((....))..)))))..))))....((...(((((...((((........))))....))))).)).------.........)))))))))). ( -25.60) >DroSec_CAF1 262 114 - 1 CCAUAGCUCUGGAGAUCACUCUGUACAACGUGAGUGCACAGAAGGUGGAGAGCCAGGUAUAUUAUAUUGUAUAAUUUUAAUGUUUAUGCAAGGA------AUUAUUGUAUUUUUAGAGUG .....(((((((((((((((((((...((....))..)))))..))))....((..(((((..((((((........)))))).)))))..)).------.........)))))))))). ( -29.60) >DroSim_CAF1 264 114 - 1 CCAUAGCUCUGGAGAUCACUCUGUACAACGUGAGUGCACAGAAGGUGGAGAGCCAGGUAUAUUAUAUUGUAUAAUUUUAAUGUUUACGUAAGUA------AUUAUUGUAUUUUUAGAGUG .....(((((((((((.(((((((...((....))..))))).(((.....)))..(((((......)))))......((((.((((....)))------).))))))))))))))))). ( -28.30) >DroEre_CAF1 306 119 - 1 CCAUAGCUCUGGAAAUCACUUUGUACAACGUGAGUGCACAGAAGGUGGAGAGCCAGGUAUGUUUUAUUGUAUUAUUACAAUUUUGAUUUAAGGAAAUACGAGUAUUGU-GUUUCAGAGUG .....(((((..((((((.(((((...((....))..))))).(((.....)))...........((((((....))))))..))))))...(((((((((...))))-)))))))))). ( -27.30) >consensus CCAUAGCUCUGGAGAUCACUCUGUACAACGUGAGUGCACAGAAGGUGGAGAGCCAGGUAUAUUAUAUUGUAUAAUUUCAAUGUUUAUGUAAGGA______AUUAUUGUAUUUUUAGAGUG .....(((((((((((.(((((((...((....))..))))).(((.....)))..(((((......)))))..................................))))))))))))). (-22.80 = -22.55 + -0.25)

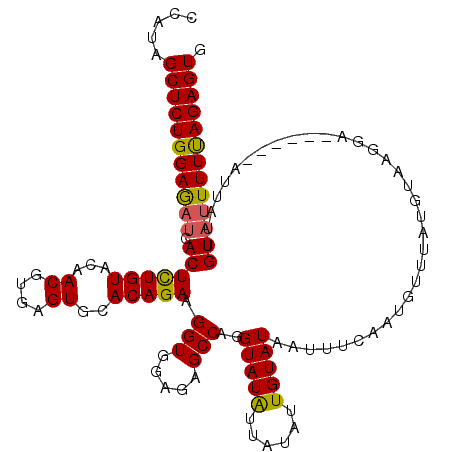

| Location | 9,601,655 – 9,601,775 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.67 |
| Mean single sequence MFE | -22.50 |
| Consensus MFE | -18.43 |
| Energy contribution | -19.48 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.737395 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9601655 120 + 23771897 UUGAAAUUAUACAAUAUAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGGUCACAAAGUAGACCAUAAUCAUGCCCAUCCACACCCAGCC ........................((((((((.(((..((((((((.......)))))))))))......)))))(((((((........)))))))..................))).. ( -26.10) >DroSec_CAF1 296 120 + 1 UUAAAAUUAUACAAUAUAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGGUCACAAAGUAGACCAUAAUCAUGCCCAUCCACACCCAGCC ........................((((((((.(((..((((((((.......)))))))))))......)))))(((((((........)))))))..................))).. ( -26.10) >DroSim_CAF1 298 120 + 1 UUAAAAUUAUACAAUAUAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGGUCACAAAGUAGACCAUAAUCAUGCCCAUCCACACCCAUCC ..........................(((....(((..((((((((.......))))))))))).......((..(((((((........))))))).))..)))............... ( -25.70) >DroEre_CAF1 345 120 + 1 UUGUAAUAAUACAAUAAAACAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAAAGUGAUUUCCAGAGCUAUGGUCACAAAGUAGACCAUAAUCAUGCCCAUCCACACCCAACC (((((....)))))............(((....(((.((.((((((.......))))))))))).......((..(((((((........))))))).))..)))............... ( -22.10) >DroYak_CAF1 297 120 + 1 UUAUAAUAAUACAAUACAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGGUCACAAAGUAAACCAUAAUCAUGCCCAUCCACACCCAGCC ..............(((.....((((((((((.(((..((((((((.......)))))))))))......))))))).)))......))).............................. ( -21.50) >DroPer_CAF1 280 95 + 1 -------------------------UGGGUCUCCACUUUCUGCGCGCUUACAUUGUACAAGCUGAUCUCCAGAGUUAUUGUCACAAAGUACACCAUAAUCAUGCCCAUCCAUACCCAGCC -------------------------(((((..................(((.((((((((.(((.....))).....)))).)))).)))..........(((......))))))))... ( -13.50) >consensus UUAAAAUUAUACAAUAUAAUAUACCUGGCUCUCCACCUUCUGUGCACUCACGUUGUACAGAGUGAUCUCCAGAGCUAUGGUCACAAAGUAGACCAUAAUCAUGCCCAUCCACACCCAGCC ..........................(((....(((..((((((((.......))))))))))).......((..(((((((........))))))).))..)))............... (-18.43 = -19.48 + 1.06)

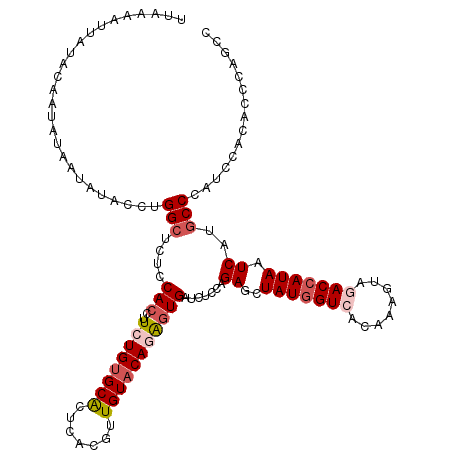
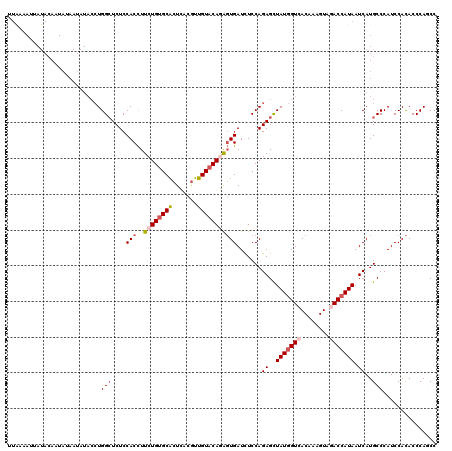
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:53:32 2006