
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,579,251 – 9,579,366 |
| Length | 115 |
| Max. P | 0.784127 |

| Location | 9,579,251 – 9,579,366 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.93 |
| Mean single sequence MFE | -35.05 |
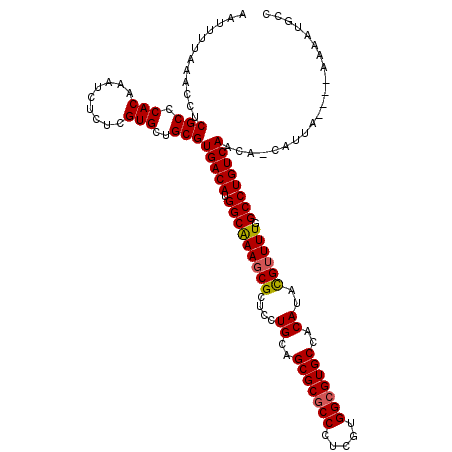
| Consensus MFE | -31.10 |
| Energy contribution | -30.93 |
| Covariance contribution | -0.17 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.784127 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
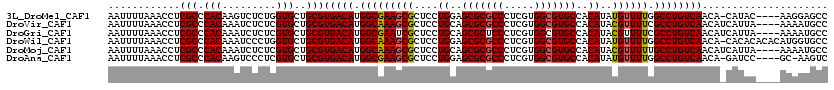
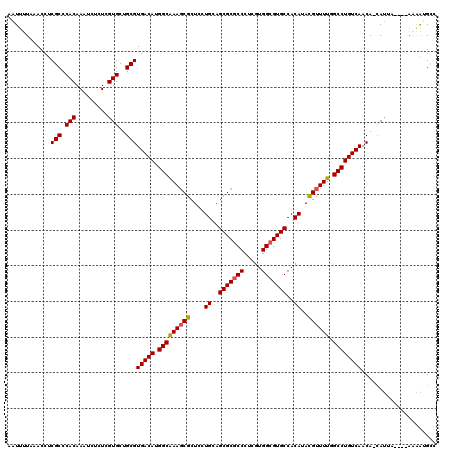
>3L_DroMel_CAF1 9579251 115 - 23771897 AAUUUUAAACCUCGCCCACAAGUCUCUGGUGCUGCGUGACAUGGCGAAGCGCUCCUGGAGCGCGCCCUCGUGGCGUGCCACAUAUGUUUUGGCCUGUCAACA-CAUAC----AAGGAGCC ...........((((((((.(((.......)))..)))....)))))...(((((((..(((((((.....)))))))..).(((((.((((....)))).)-)))).----.)))))). ( -39.90) >DroVir_CAF1 803 116 - 1 AAUUUUAAACCUCGCCCACAAAUCUCUCGUGCUGCGUGACAUGGCAAAGCGCUCCUGCAGCGCGCCCUCGUGGCGUGCCACAUACGUUUUCGCCUGUCAACAUCAUUA----AAAAUGCC ..((((((....(((.(((.........)))..)))(((((.((((((((((....)).(((((((.....))))))).......))))).))))))))......)))----)))..... ( -34.50) >DroGri_CAF1 1146 116 - 1 AAUUUUAAACCUCGCCCACAAAUCUCUCGUGCUGCGUGACAUGGCGAAUCGCUCCUGCAGCGCUCCCUCGUGGCGUGCCACAUACGUUUUCGCCUGUCAACAUCAUUA----AAAAUGCC ..((((((....(((.(((.........)))..)))(((((.((((((.((((.....)))).......((((....)))).......)))))))))))......)))----)))..... ( -30.20) >DroWil_CAF1 2012 119 - 1 AAUUUUAAACCUCGCCCACAAAUCCCUGGUGCUGCGUGACAUGGCAAAGCGCUCCUGGAGCGCGCCCUCGUGGCGUGCCACAUAUGUUUUGGCCUGUCAACA-CACACACACAUGGUGCC .............(((((((......))(((.((.((((((.(((((((((....((..(((((((.....)))))))..))..)))))).))))))).)).-))))).....))).)). ( -37.80) >DroMoj_CAF1 1194 116 - 1 AAUUUUAAACCUCGCCCACAAAUCUCUCGUGCUGCGUGACAUGGCAAAGCGCUCCUGCAGCGCGCCCUCGUGGCGUGCCACAUACGUUUUUGCCUGUCAACAUCAUUA----AAAAUGCC ..((((((....(((.(((.........)))..)))(((((.((((((((((....)).(((((((.....))))))).......).))))))))))))......)))----)))..... ( -34.50) >DroAna_CAF1 4717 114 - 1 AAUUUUAAACCUCGCCCACAAGUCCCUCGUGCUGCGUGACAUGGCGAAGCGCUCCUGGAGCGCGCCCUCGUGGCGUGCCACAUAUGUUUUGGCCUGUCAACA-GAUCC----GC-AAGUC .............((.(((.((...)).)))(((..(((((.(((((((((....((..(((((((.....)))))))..))..)))))).)))))))).))-)....----))-..... ( -33.40) >consensus AAUUUUAAACCUCGCCCACAAAUCUCUCGUGCUGCGUGACAUGGCAAAGCGCUCCUGCAGCGCGCCCUCGUGGCGUGCCACAUACGUUUUGGCCUGUCAACA_CAUUA____AAAAUGCC ............(((.(((.........)))..)))(((((.(((((((((....((..(((((((.....)))))))..))..)))))).))))))))..................... (-31.10 = -30.93 + -0.17)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:53:25 2006