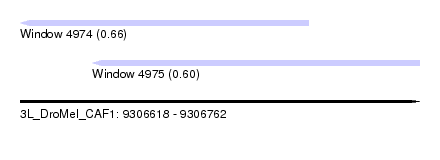

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,306,618 – 9,306,762 |
| Length | 144 |
| Max. P | 0.659450 |

| Location | 9,306,618 – 9,306,722 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 67.45 |
| Mean single sequence MFE | -37.73 |
| Consensus MFE | -20.41 |
| Energy contribution | -19.48 |
| Covariance contribution | -0.94 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.659450 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
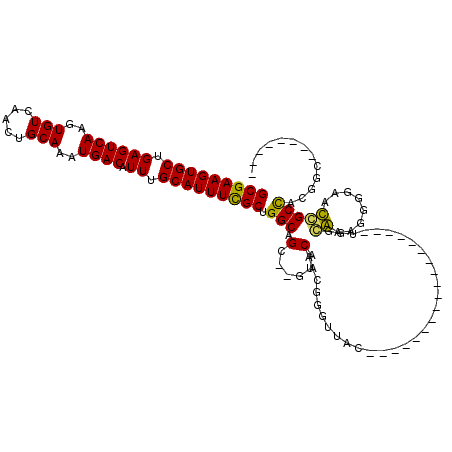
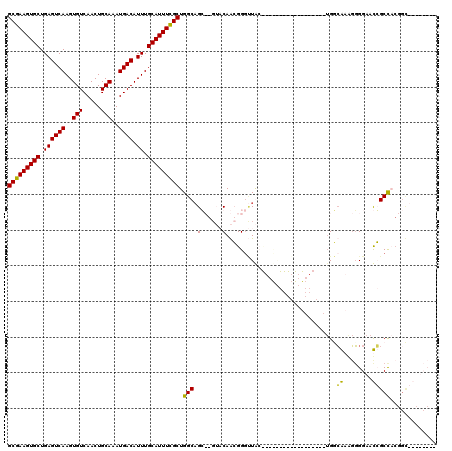
>3L_DroMel_CAF1 9306618 104 - 23771897 GCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUCGCUGGCAGG--GAUCAAGGGGCUGAUGACGGGGGGAGG------GAGGCUCUGGAACCGCUCCGGU-------- (((((((((.((((((..(((.....)))..)))).)).)))))))))(((.(((--(.((.((((.((..(..(....)..).------.)).)))).)).)).)))))..-------- ( -35.20) >DroPse_CAF1 173013 102 - 1 GCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUCGCUGGCAGCCGGUACAGCGGGUUAC------------------UGGCAAAGGGGAAUGGCCACGGCUACUCCAC (((((((((.((((((..(((.....)))..)))).)).))))))))).....(((((((........)))------------------)))).....(((.((((....)))).))).. ( -40.20) >DroAna_CAF1 166718 103 - 1 GCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUUGCUGGCCGG-----CAAGGGAAUGGCGGUAGGGUGUGCCGGGGUGUCGCGGGGGAACCGCUC------------ (((((((((.((((((..(((.....)))..)))).)).)))))))))..((((.-----((......)).))))..((....))........((((.....))))..------------ ( -35.30) >DroPer_CAF1 174234 102 - 1 GCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUCGCUGGCAGCCGGUACAGCGGGUUAC------------------UGGCAAAGGGGAAUGGCCACGGCUACUCCAC (((((((((.((((((..(((.....)))..)))).)).))))))))).....(((((((........)))------------------)))).....(((.((((....)))).))).. ( -40.20) >consensus GCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUCGCUGGCAGC__GUACAACGGGUUAC__________________UGGCAAAGGGGAACCGCCACGGC________ (((((((((.((((((..(((.....)))..)))).)).))))))))).(((.(......)...............................((........)))))............. (-20.41 = -19.48 + -0.94)

| Location | 9,306,644 – 9,306,762 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.99 |
| Mean single sequence MFE | -37.45 |
| Consensus MFE | -26.56 |
| Energy contribution | -26.38 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.595398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
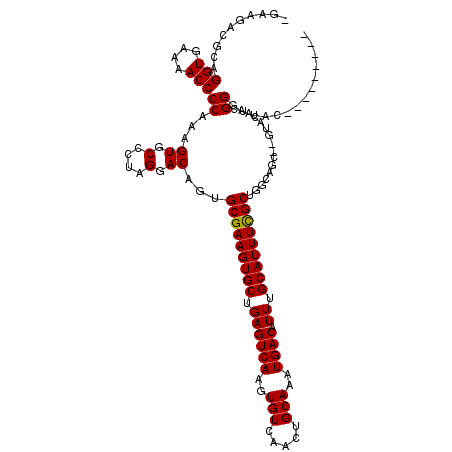
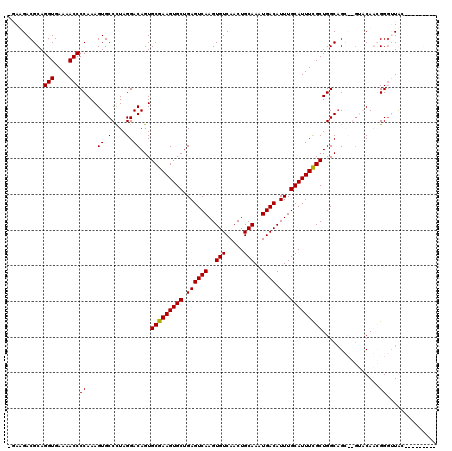
>3L_DroMel_CAF1 9306644 118 - 23771897 CAAAGACGCAGGUGAAAACCCCAAAGUGCCCUAGGACAGUGCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUCGCUGGCAGG--GAUCAAGGGGCUGAUGACGGGGG ..................((((.....(((((..(((.(((((((((((.((((((..(((.....)))..)))).)).)))))))))..))...--).))..))))).......)))). ( -37.50) >DroPse_CAF1 173044 110 - 1 -GAUGACGCAGGUGAAAACCCCAAAGUGCCUUAGGACAGUGCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUCGCUGGCAGCCGGUACAGCGGGUUAC--------- -.....(((.(((....))).....(((((...((.....(((((((((.((((((..(((.....)))..)))).)).)))))))))......))))))).)))......--------- ( -37.70) >DroAna_CAF1 166746 113 - 1 --AAGACGCAGGUGAAAACCCCAAAGUGCCCUAGGACAGUGCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUUGCUGGCCGG-----CAAGGGAAUGGCGGUAGGGU --....(((.(((....)))((....((((...((.(...(((((((((.((((((..(((.....)))..)))).)).)))))))))..)))))-----))..))....)))....... ( -36.90) >DroPer_CAF1 174265 110 - 1 -GAUGACGCAGGUGAAAACCCCAAAGUGCCUUAGGACAGUGCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUCGCUGGCAGCCGGUACAGCGGGUUAC--------- -.....(((.(((....))).....(((((...((.....(((((((((.((((((..(((.....)))..)))).)).)))))))))......))))))).)))......--------- ( -37.70) >consensus _GAAGACGCAGGUGAAAACCCCAAAGUGCCCUAGGACAGUGCGAAGUGCUGAGUCAAGUGUCAACUGCAAAUGACAUUUGCAUUUCGCUGGCAGC__GUACAACGGGUUAC_________ ..........(((....)))((...((.(....).))...(((((((((.((((((..(((.....)))..)))).)).)))))))))................)).............. (-26.56 = -26.38 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:46 2006