
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,259,969 – 9,260,066 |
| Length | 97 |
| Max. P | 0.979796 |

| Location | 9,259,969 – 9,260,066 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 82.20 |
| Mean single sequence MFE | -19.66 |
| Consensus MFE | -10.30 |
| Energy contribution | -13.00 |
| Covariance contribution | 2.70 |
| Combinations/Pair | 1.18 |
| Mean z-score | -3.26 |
| Structure conservation index | 0.52 |
| SVM decision value | 1.85 |
| SVM RNA-class probability | 0.979796 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9259969 97 + 23771897 AAGUAAUUUGAAAAUUAAAUUUCAAUUUCGAUUUCCAAAGUGAAAAUGCGAGGAAAAUGCGUUGAAAAAUGCAUAGCUCGCGUUGAGAAUAAAAUGG .......((((((......))))))((((...(((......)))((((((((....(((((((....)))))))..))))))))))))......... ( -22.00) >DroSec_CAF1 106746 89 + 1 AAGUAAUUUGAAAAUUAAAC--------CGAUUUCAAAAGUGAAAAUGCGAUGAAAAUGCGUUGAAAAAUGAAUAGCUCGCGUUGAGAACAAAAUUG ......(((((((.......--------...))))))).((...(((((((.(......((((....)))).....))))))))....))....... ( -15.00) >DroSim_CAF1 106997 88 + 1 AAGUAAUUUGAAAAUUAAAC--------CGUUUUCAAAAGUGAAAAUGCGAGG-AAAUGCGCUGAAAAAUGCAUGGCUCGCGUUGAGAACAAAAUUG ......(((((((((.....--------.))))))))).((...((((((((.-..(((((.(....).)))))..))))))))....))....... ( -23.60) >DroEre_CAF1 129526 88 + 1 AAGUAGUUUGAAAAU--AAG-------CAGAUUUUAAAAGUGAAAAUGCGAGAAAAAUGCGUUGAAAAAUGCAUAGCGCACUUUCAGAGAGAAAUGG ....((((((.....--...-------)))))).......(((((.((((......(((((((....)))))))..)))).)))))........... ( -18.20) >DroYak_CAF1 124752 89 + 1 AAGUAAUUUGAAAAUUAAAG-------CAAAU-GCAAAAGUGAAAAUGCGCGAAAAAUGCGUUGAAAAAUGCAUAGCGCACUUUGGGAGCAAAAUUG ......((((....((((((-------(....-))...........(((((.....(((((((....))))))).))))).)))))...)))).... ( -19.50) >consensus AAGUAAUUUGAAAAUUAAAC________CGAUUUCAAAAGUGAAAAUGCGAGGAAAAUGCGUUGAAAAAUGCAUAGCUCGCGUUGAGAACAAAAUUG .......................................((...((((((((....(((((((....)))))))..))))))))....))....... (-10.30 = -13.00 + 2.70)



| Location | 9,259,969 – 9,260,066 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 82.20 |
| Mean single sequence MFE | -13.37 |
| Consensus MFE | -7.50 |
| Energy contribution | -7.62 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.875621 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9259969 97 - 23771897 CCAUUUUAUUCUCAACGCGAGCUAUGCAUUUUUCAACGCAUUUUCCUCGCAUUUUCACUUUGGAAAUCGAAAUUGAAAUUUAAUUUUCAAAUUACUU ........(((.....(((((..((((..........))))....)))))((((((.....)))))).))).((((((......))))))....... ( -17.00) >DroSec_CAF1 106746 89 - 1 CAAUUUUGUUCUCAACGCGAGCUAUUCAUUUUUCAACGCAUUUUCAUCGCAUUUUCACUUUUGAAAUCG--------GUUUAAUUUUCAAAUUACUU ................((((..........................)))).........(((((((...--------.......)))))))...... ( -6.77) >DroSim_CAF1 106997 88 - 1 CAAUUUUGUUCUCAACGCGAGCCAUGCAUUUUUCAGCGCAUUU-CCUCGCAUUUUCACUUUUGAAAACG--------GUUUAAUUUUCAAAUUACUU ................(((((..((((..........))))..-.))))).........((((((((..--------......))))))))...... ( -14.70) >DroEre_CAF1 129526 88 - 1 CCAUUUCUCUCUGAAAGUGCGCUAUGCAUUUUUCAACGCAUUUUUCUCGCAUUUUCACUUUUAAAAUCUG-------CUU--AUUUUCAAACUACUU ...........((((((((((..((((..........))))......)))))))))).............-------...--............... ( -13.60) >DroYak_CAF1 124752 89 - 1 CAAUUUUGCUCCCAAAGUGCGCUAUGCAUUUUUCAACGCAUUUUUCGCGCAUUUUCACUUUUGC-AUUUG-------CUUUAAUUUUCAAAUUACUU (((...(((....(((((((((.((((..........)))).....))))))))).......))-).)))-------.................... ( -14.80) >consensus CAAUUUUGUUCUCAACGCGAGCUAUGCAUUUUUCAACGCAUUUUCCUCGCAUUUUCACUUUUGAAAUCG________CUUUAAUUUUCAAAUUACUU ................(((((..((((..........))))....)))))............................................... ( -7.50 = -7.62 + 0.12)

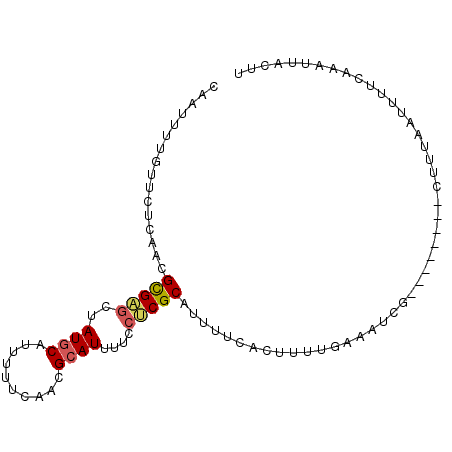

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:27 2006