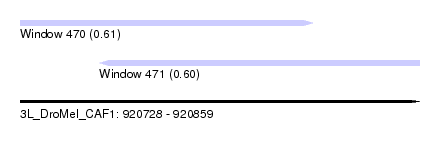

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 920,728 – 920,859 |
| Length | 131 |
| Max. P | 0.610640 |

| Location | 920,728 – 920,824 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 84.78 |
| Mean single sequence MFE | -28.74 |
| Consensus MFE | -19.40 |
| Energy contribution | -19.24 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.610640 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
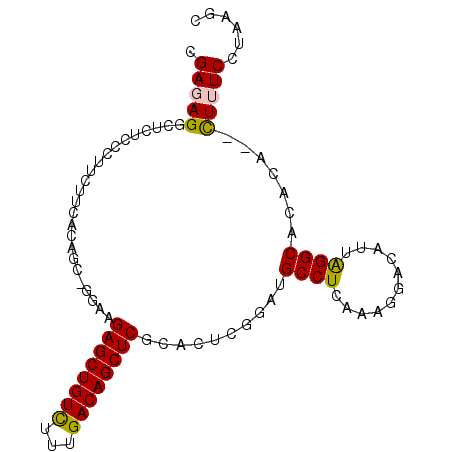
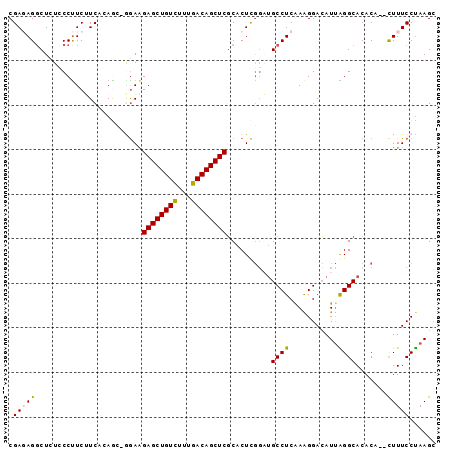
>3L_DroMel_CAF1 920728 96 + 23771897 CGAGAGUCUCUCCCUUCUUCACAACUGAAAGAGCUGUUUUUGACAGCUCGCACUUGGAUGCCUCAAAGGACAUCAGGCACACAAAUUUUCCUAAGC .((((((.....(((......(((.((...((((((((...)))))))).)).)))((((((.....)).))))))).......))))))...... ( -24.10) >DroSec_CAF1 55758 92 + 1 CGAUAGGCUCUCCCUUCUUCAGAG--AGAAGAGCUGUCUGUGACAGCUCGCACUCGGAUGCCUCAAAGGACAUUAGGCACACA--CUUUCCUAAGC ...(((((((((.........)))--))..((((((((...))))))))((.((..((((((.....)).)))))))).....--....))))... ( -27.30) >DroSim_CAF1 48751 92 + 1 CGAUAGGCUCUCCCUUCUUCAGAG--AGAAGAGCUGUCUUUGACAGCUCCCACUCAUAUGCCUCAAGGGAGAUUAGGCACACA--CUUUCCUAAGC ......((((((((((...((..(--((..((((((((...))))))))...)))...))....))))))))(((((......--....))))))) ( -29.10) >DroEre_CAF1 59105 94 + 1 CGAGAGGCUAUCCCCUCAUCACAGCUGGAAGAGCUGUCUUUGACAGCUCGCACUCGGUUGCCUCGAAGGACAUUGGGCACACA--CUUUCCCAAGC .((((((......)))).))...(((((((((((((((...))))))))......(..((((.(((......)))))))..).--..))))..))) ( -30.70) >DroYak_CAF1 58372 94 + 1 CGAGAGGCUCUGCCUUCAUCACAACGGGAGGAGCUGUCUUUGACAGCUCGCACUCGGUGGCCUCAAAGGACAUUGGGCACACA--CUUUCCUAAGG .(((((((...)))))..))....((((.(((((((((...)))))))).).))))((((((.(((......)))))).))).--........... ( -32.50) >consensus CGAGAGGCUCUCCCUUCUUCACAGC_GGAAGAGCUGUCUUUGACAGCUCGCACUCGGAUGCCUCAAAGGACAUUAGGCACACA__CUUUCCUAAGC .(((((........................((((((((...))))))))..........((((...........)))).......)))))...... (-19.40 = -19.24 + -0.16)

| Location | 920,754 – 920,859 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 82.29 |
| Mean single sequence MFE | -31.42 |
| Consensus MFE | -19.20 |
| Energy contribution | -20.36 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.600811 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
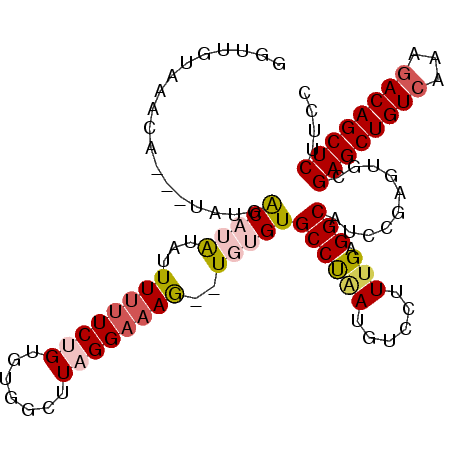
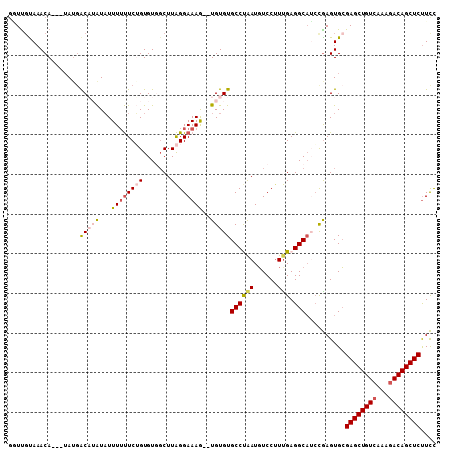
>3L_DroMel_CAF1 920754 105 - 23771897 GGUUGUAAACA---UAUGACAUAUAUUUUUUCUGUGUUGCUUAGGAAAAUUUGUGUGCCUGAUGUCCUUUGAGGCAUCCAAGUGCGAGCUGUCAAAAACAGCUCUUUC .((..((....---))..)).......(((((((.......)))))))(((((..(((((.(.......).)))))..)))))..(((((((.....))))))).... ( -28.70) >DroSec_CAF1 55782 103 - 1 UGUUGCAAACA---UAGGGCAUAUAUUUUUUCUGUGUGGCUUAGGAAAG--UGUGUGCCUAAUGUCCUUUGAGGCAUCCGAGUGCGAGCUGUCACAGACAGCUCUUCU ....(((.(((---(.((((((((((((((.(((.......))))))))--))))))))).)))).(((.((....)).))))))((((((((...)))))))).... ( -37.10) >DroSim_CAF1 48775 103 - 1 GAUUGUAAACA---GAUGACUUAUAUUUUUUCUGUGUGGCUUAGGAAAG--UGUGUGCCUAAUCUCCCUUGAGGCAUAUGAGUGGGAGCUGUCAAAGACAGCUCUUCU ...........---....((((....((((((((.......))))))))--.((((((((...........))))))))))))((((((((((...)))))))))).. ( -28.30) >DroEre_CAF1 59131 103 - 1 GCUCGCAAACA---UAUGACUAGUUUUUUCUCGGUGUGGCUUGGGAAAG--UGUGUGCCCAAUGUCCUUCGAGGCAACCGAGUGCGAGCUGUCAAAGACAGCUCUUCC ....((((((.---........)))....((((((.((.(((((((...--((......))...)))..)))).)))))))))))((((((((...)))))))).... ( -32.60) >DroYak_CAF1 58398 105 - 1 GUUUCUAAACAAAGUAUCACAAAUAUUU-UUCGGUGUACCUUAGGAAAG--UGUGUGCCCAAUGUCCUUUGAGGCCACCGAGUGCGAGCUGUCAAAGACAGCUCCUCC ..........(((((((.....))))))-)((((((..((((((((...--((......))...))))..)))).))))))....((((((((...)))))))).... ( -30.40) >consensus GGUUGUAAACA___UAUGACAUAUAUUUUUUCUGUGUGGCUUAGGAAAG__UGUGUGCCUAAUGUCCUUUGAGGCAUCCGAGUGCGAGCUGUCAAAGACAGCUCUUCC ..................(((((...((((((((.......))))))))..)))))((((((......))).)))..........((((((((...)))))))).... (-19.20 = -20.36 + 1.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:39:02 2006