
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 9,117,611 – 9,117,707 |
| Length | 96 |
| Max. P | 0.853712 |

| Location | 9,117,611 – 9,117,707 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.04 |
| Mean single sequence MFE | -28.17 |
| Consensus MFE | -22.43 |
| Energy contribution | -23.27 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.669601 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9117611 96 + 23771897 GGUAUGGCACAAAUAAUUGUUACCGCUGCAACCCCCAUGGUUGAAGACGGUGCCGAUAAAUCUUUCCCACCAGUAGGCGAGAGCGAUGCACAGCCA------------------------ (((.((((((.....((((.(((((((.(((((.....))))).)).))))).))))....(((((((.......)).))))).).))).))))).------------------------ ( -28.90) >DroVir_CAF1 5604 90 + 1 GGUAUGGCACAAAUAAUUGUUACCGCUGCAACCCCAAUGGUUGAAGACGGUGCCGAUAAAUCUUUCCCACCAGUAGGAGAAAGGGACAAU------------------------------ ....((((((((....)))).((((((.(((((.....))))).)).)))))))).....((((((((.......)).))))))......------------------------------ ( -25.00) >DroGri_CAF1 4761 93 + 1 GGUAUGGCACAAAUAAUUGUUACCGCUGCAACCCCAAUGGUUGAAGACGGUGCCGAUAAAUCUUUCCCACCAGUAGGAGAGAGCGACAAU---UCA------------------------ ......((.......((((.(((((((.(((((.....))))).)).))))).))))...(((((((........)))))))))......---...------------------------ ( -26.90) >DroWil_CAF1 6221 96 + 1 GGUAUGGCACAAAUAAUAGUUACCGCUGCAACCCCAAUGGUUGAAGACGGUGCAGAUAAAUCUUUCCCACCAGUAGGAGAGAAGGAGGAACAGCAA------------------------ ......((..........((.((((((.(((((.....))))).)).)))))).......(((((((........)))))))..........))..------------------------ ( -24.30) >DroMoj_CAF1 4842 93 + 1 GGUAUGGCACAAAUAAUUGUUACCGCUGCAACCCCAAUGGUUGAAGACGGUGCCGAUAAAUCUUUCCCACCAGUAGGAGAGAGCGACAAU---ACA------------------------ .((((.((.......((((.(((((((.(((((.....))))).)).))))).))))...(((((((........)))))))))....))---)).------------------------ ( -28.60) >DroAna_CAF1 5821 120 + 1 GGUAUGGCACAAAUAAUAGUUACCGCUGCAACCCCCAUGGUUGAAGACGGUGCCGAUAAAUCUUUCCCACCAGUAGGAGAGAGCGAUUCCCAGUCCGAUAAGGACACGGAUACGGAUAAG ((.((.((..........(.(((((((.(((((.....))))).)).))))).)......(((((((........))))))))).)).))..((((.....)))).((....))...... ( -35.30) >consensus GGUAUGGCACAAAUAAUUGUUACCGCUGCAACCCCAAUGGUUGAAGACGGUGCCGAUAAAUCUUUCCCACCAGUAGGAGAGAGCGACAAU___CCA________________________ ....((((((((....)))).((((((.(((((.....))))).)).)))))))).....(((((((........)))))))...................................... (-22.43 = -23.27 + 0.83)



| Location | 9,117,611 – 9,117,707 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.04 |
| Mean single sequence MFE | -30.80 |
| Consensus MFE | -25.42 |
| Energy contribution | -25.78 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853712 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 9117611 96 - 23771897 ------------------------UGGCUGUGCAUCGCUCUCGCCUACUGGUGGGAAAGAUUUAUCGGCACCGUCUUCAACCAUGGGGGUUGCAGCGGUAACAAUUAUUUGUGCCAUACC ------------------------..((((((.(((..(((((((....)))))))..))).)).))))((((.((.(((((.....))))).)))))).((((....))))........ ( -33.00) >DroVir_CAF1 5604 90 - 1 ------------------------------AUUGUCCCUUUCUCCUACUGGUGGGAAAGAUUUAUCGGCACCGUCUUCAACCAUUGGGGUUGCAGCGGUAACAAUUAUUUGUGCCAUACC ------------------------------.......((((((((.....).))))))).......(((((((.((.(((((.....))))).)))))).((((....)))))))..... ( -27.30) >DroGri_CAF1 4761 93 - 1 ------------------------UGA---AUUGUCGCUCUCUCCUACUGGUGGGAAAGAUUUAUCGGCACCGUCUUCAACCAUUGGGGUUGCAGCGGUAACAAUUAUUUGUGCCAUACC ------------------------.((---...(((..((((.((....)).))))..)))...))(((((((.((.(((((.....))))).)))))).((((....)))))))..... ( -30.60) >DroWil_CAF1 6221 96 - 1 ------------------------UUGCUGUUCCUCCUUCUCUCCUACUGGUGGGAAAGAUUUAUCUGCACCGUCUUCAACCAUUGGGGUUGCAGCGGUAACUAUUAUUUGUGCCAUACC ------------------------..(((((..(((((((((.((....)).)))))(((....)))..................))))..)))))((((.(........)))))..... ( -25.90) >DroMoj_CAF1 4842 93 - 1 ------------------------UGU---AUUGUCGCUCUCUCCUACUGGUGGGAAAGAUUUAUCGGCACCGUCUUCAACCAUUGGGGUUGCAGCGGUAACAAUUAUUUGUGCCAUACC ------------------------.((---((.(((..((((.((....)).))))..))).....(((((((.((.(((((.....))))).)))))).((((....))))))))))). ( -29.70) >DroAna_CAF1 5821 120 - 1 CUUAUCCGUAUCCGUGUCCUUAUCGGACUGGGAAUCGCUCUCUCCUACUGGUGGGAAAGAUUUAUCGGCACCGUCUUCAACCAUGGGGGUUGCAGCGGUAACUAUUAUUUGUGCCAUACC .......(((((((.((((.....)))))))(((((..((((.((....)).))))..)))))...(((((((.((.(((((.....))))).)))))).((........))))))))). ( -38.30) >consensus ________________________UGA___AUUGUCGCUCUCUCCUACUGGUGGGAAAGAUUUAUCGGCACCGUCUUCAACCAUUGGGGUUGCAGCGGUAACAAUUAUUUGUGCCAUACC .................................(((..((((.((....)).))))..))).....(((((((.((.(((((.....))))).)))))).((((....)))))))..... (-25.42 = -25.78 + 0.36)

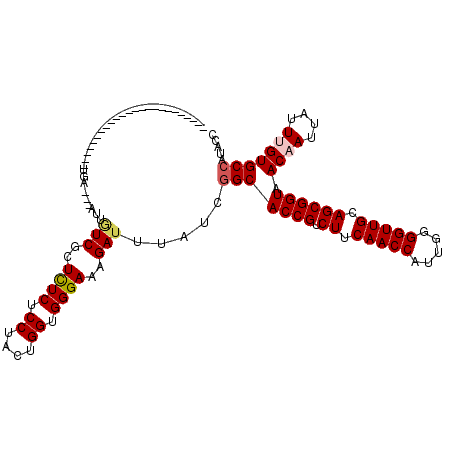

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:50:14 2006