
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 8,545,747 – 8,545,863 |
| Length | 116 |
| Max. P | 0.991299 |

| Location | 8,545,747 – 8,545,851 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.58 |
| Mean single sequence MFE | -34.10 |
| Consensus MFE | -34.10 |
| Energy contribution | -33.35 |
| Covariance contribution | -0.75 |
| Combinations/Pair | 1.20 |
| Mean z-score | -5.47 |
| Structure conservation index | 1.00 |
| SVM decision value | 2.26 |
| SVM RNA-class probability | 0.991299 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 8545747 104 + 23771897 AAC---------------U-AAUUGAUGGUUCCAGUGAGAUAUGUUUGAUAUUCUUGGUUGUUUCAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGACCCUCGCCAAUCU ...---------------.-....((.(((..((((((((((((((((((..((((((.((...(........)..)).)))))))))))))))))).))))))..))).))........ ( -34.10) >DroVir_CAF1 825 115 + 1 AACUACC----CCCCA-CUCAUCUAAUGAUUCCAGUAAGAUAUGUUUGAUAUUCUUGGUUGUUACAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUUGUUGAAACUCU .......----.....-......(((..((..((((((((((((((((((..((((((.((..((........)).)).)))))))))))))))))).))))))..))..)))....... ( -32.50) >DroPse_CAF1 926 104 + 1 AAC---------------U-AAUUGGGGAUUCCAGUGAGAUAUGUUUGAUAUUCUUGGUUGUUACAUUCAAGAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUACCCGGAAAUCU ...---------------.-..((.(((((..((((((((((((((((((..((((((.((..((........)).)).)))))))))))))))))).))))))..)).))).))..... ( -35.90) >DroGri_CAF1 912 115 + 1 ACCAACU----ACCCA-CUCAUCUAAUGAUUCCAGUGAGAUAUGUUUGAUAUUCUUGGUUGUUACAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUUGUUGAAACUCU .......----.....-......(((..((..((((((((((((((((((..((((((.((..((........)).)).)))))))))))))))))).))))))..))..)))....... ( -32.60) >DroMoj_CAF1 851 120 + 1 AACUACCUAUCCACUACCUCAUCUAACGAUUCCAGUGAGAUAUGUUUGAUAUUCUUGGUUGUUACAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUCGCUUAAACUCU .......................(((((((..((((((((((((((((((..((((((.((..((........)).)).)))))))))))))))))).))))))..)))).)))...... ( -33.60) >DroPer_CAF1 932 104 + 1 AAC---------------U-AAUUGGGGAUUCCAGUGAGAUAUGUUUGAUAUUCUUGGUUGUUACAUUCAAGAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUACCCGGAAAUCU ...---------------.-..((.(((((..((((((((((((((((((..((((((.((..((........)).)).)))))))))))))))))).))))))..)).))).))..... ( -35.90) >consensus AAC_______________U_AACUAAUGAUUCCAGUGAGAUAUGUUUGAUAUUCUUGGUUGUUACAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUACCCGAAAAUCU .......................(((((((..((((((((((((((((((..((((((.((..((........)).)).)))))))))))))))))).))))))..)))))))....... (-34.10 = -33.35 + -0.75)



| Location | 8,545,747 – 8,545,851 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.58 |
| Mean single sequence MFE | -26.18 |
| Consensus MFE | -20.38 |
| Energy contribution | -19.30 |
| Covariance contribution | -1.08 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.785860 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 8545747 104 - 23771897 AGAUUGGCGAGGGUCACAGUAAUAAUAUGUUUGAUUCCUGGGUGAACUUUUGAAUGAAACAACCAAGAAUAUCAAACAUAUCUCACUGGAACCAUCAAUU-A---------------GUU .((((((....(((..((((....(((((((((((..((.(((....((((....))))..))).))...)))))))))))...))))..))).))))))-.---------------... ( -26.90) >DroVir_CAF1 825 115 - 1 AGAGUUUCAACAAUCACAGUAAUAAUAUGUUUGAUUCCUGGGUGAACUUUUGAAUGUAACAACCAAGAAUAUCAAACAUAUCUUACUGGAAUCAUUAGAUGAG-UGGGG----GGUAGUU .....(((..((.((.((((((..(((((((((((..((.(((..((........))....))).))...))))))))))).))))))..(((....))))).-))..)----))..... ( -24.80) >DroPse_CAF1 926 104 - 1 AGAUUUCCGGGUAUCACAGUAAUAAUAUGUUUGAUUCCUGGGUGAACUCUUGAAUGUAACAACCAAGAAUAUCAAACAUAUCUCACUGGAAUCCCCAAUU-A---------------GUU ........(((.((..((((....(((((((((((..((.(((..((........))....))).))...)))))))))))...))))..)).)))....-.---------------... ( -25.90) >DroGri_CAF1 912 115 - 1 AGAGUUUCAACAAUCACAGUAAUAAUAUGUUUGAUUCCUGGGUGAACUUUUGAAUGUAACAACCAAGAAUAUCAAACAUAUCUCACUGGAAUCAUUAGAUGAG-UGGGU----AGUUGGU ......(((((.............(((((((((((..((.(((..((........))....))).))...)))))))))))((((((...(((....))).))-)))).----.))))). ( -26.00) >DroMoj_CAF1 851 120 - 1 AGAGUUUAAGCGAUCACAGUAAUAAUAUGUUUGAUUCCUGGGUGAACUUUUGAAUGUAACAACCAAGAAUAUCAAACAUAUCUCACUGGAAUCGUUAGAUGAGGUAGUGGAUAGGUAGUU ...((((((.((((..((((....(((((((((((..((.(((..((........))....))).))...)))))))))))...))))..)))))))))).................... ( -27.60) >DroPer_CAF1 932 104 - 1 AGAUUUCCGGGUAUCACAGUAAUAAUAUGUUUGAUUCCUGGGUGAACUCUUGAAUGUAACAACCAAGAAUAUCAAACAUAUCUCACUGGAAUCCCCAAUU-A---------------GUU ........(((.((..((((....(((((((((((..((.(((..((........))....))).))...)))))))))))...))))..)).)))....-.---------------... ( -25.90) >consensus AGAGUUUCAACAAUCACAGUAAUAAUAUGUUUGAUUCCUGGGUGAACUUUUGAAUGUAACAACCAAGAAUAUCAAACAUAUCUCACUGGAAUCAUCAAAU_A_______________GUU ........((..((..((((....(((((((((((..((.(((..((........))....))).))...)))))))))))...))))..))..))........................ (-20.38 = -19.30 + -1.08)
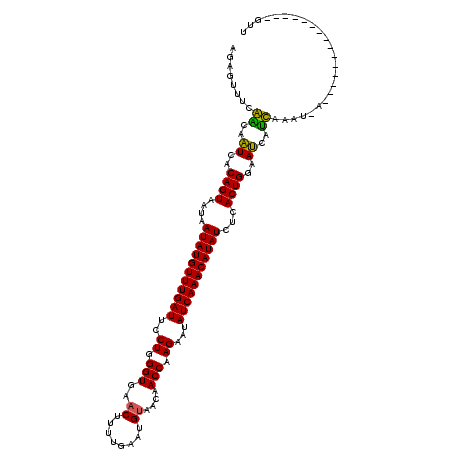



| Location | 8,545,771 – 8,545,863 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 86.31 |
| Mean single sequence MFE | -18.66 |
| Consensus MFE | -16.88 |
| Energy contribution | -17.05 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.09 |
| SVM RNA-class probability | 0.987663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 8545771 92 + 23771897 UAUGUUUGAUAUUCUUGGUUGUUUCAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGACCCUCGCCAAUCUACUAAAA-CACUC------- ((((((((((..((((((.((...(........)..)).))))))))))))))))........(((.....))).............-.....------- ( -18.10) >DroVir_CAF1 860 99 + 1 UAUGUUUGAUAUUCUUGGUUGUUACAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUUGUUGAAACUCUACUAAA-CCAACCAAAACCA ((((((((((..((((((.((..((........)).)).))))))))))))))))........((....((((.............-.))))....)).. ( -18.94) >DroPse_CAF1 950 97 + 1 UAUGUUUGAUAUUCUUGGUUGUUACAUUCAAGAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUACCCGGAAAUCUACUAAAC-CCCCA-AACCC- ((((((((((..((((((.((..((........)).)).))))))))))))))))......(((.(....)))).............-.....-.....- ( -18.50) >DroWil_CAF1 905 94 + 1 UAUGUUUGAUAUUCUUGGUUGUUACAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUCGUCGG-AAUCUACUAAAA-ACCAA---UUC- ((((((((((..((((((.((..((........)).)).))))))))))))))))......(((.......)))-............-.....---...- ( -18.70) >DroAna_CAF1 844 92 + 1 UAUGUUUGAUAUUCUUGGUUGUUACAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGACUCUCGCCAAUCUACUAAAA-UACAC------- ((((((((((..((((((.((..((........)).)).))))))))))))))))........(((.....))).............-.....------- ( -19.20) >DroPer_CAF1 956 97 + 1 UAUGUUUGAUAUUCUUGGUUGUUACAUUCAAGAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUACCCGGAAAUCUACUAAAC-CCCCA-AACCC- ((((((((((..((((((.((..((........)).)).))))))))))))))))......(((.(....)))).............-.....-.....- ( -18.50) >consensus UAUGUUUGAUAUUCUUGGUUGUUACAUUCAAAAGUUCACCCAGGAAUCAAACAUAUUAUUACUGUGAUACUCGGAAAUCUACUAAAA_CACCA____CC_ ((((((((((..((((((.((..((........)).)).))))))))))))))))............................................. (-16.88 = -17.05 + 0.17)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:46:38 2006