
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 8,200,369 – 8,200,508 |
| Length | 139 |
| Max. P | 0.663907 |

| Location | 8,200,369 – 8,200,483 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 79.85 |
| Mean single sequence MFE | -26.29 |
| Consensus MFE | -17.77 |
| Energy contribution | -18.10 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.663907 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 8200369 114 + 23771897 UUCGGC----UCCAACGGACUGUUUAAUUUACGCGCUGCUAUUAUUUUAAUCUUUUGCGCUACACACAGCGUGAAAUGACUUUCACCUGGCCUCGAGCAGCGGCUUCAAAGAUAUAGA ((((..----.....))))((((.........((((((((..........(((((..((((......))))..))).))(........)......)))))).)).........)))). ( -23.47) >DroVir_CAF1 31055 102 + 1 UCCUAC----UCCUUCGGGCUGUUUAAUUUACGCGCUGCUAUUAUUUUAAUCUUUUGCGCAACA-ACAGCGUGAAAUGACUUUCACCUGGCCUCUUGCUUUGGGUUU----------- .((((.----......((((............((((....................))))....-.(((.((((((....))))))))))))).......))))...----------- ( -22.89) >DroPse_CAF1 28054 110 + 1 UCCAGC----UCCAACGGGCUGUUUAAUUUACGCGCUGCUAUUAUUUUAAUCUUUUGCGCUACACCCAGCGUGAAAUGACUUUCACCUGGCCCCG-GCGCCGGCCCCC-UGCUAUG-- ...(((----......((((((..........((...)).................(((((....((((.((((((....))))))))))....)-))))))))))..-.)))...-- ( -31.10) >DroMoj_CAF1 26768 106 + 1 UGCUGCUCGGUCUUUCUGGCUGUUUAAUUUACGCGCUGCUAUUAUUUUAAUCUUUUGCGCUACA-ACAGCGUGAAAUGACUUUCACCUGGCCUUUUGCUUCUGCUCU----------- .((.((..((((......((((((.....((.((((....................)))))).)-)))))((((((....))))))..))))....))....))...----------- ( -22.55) >DroAna_CAF1 26801 98 + 1 UUCGGC----UCCAACGGGCUGUUUAAUUUACGCGCAGCUAUUAUUUUAAUCUUUUGCGCUACACACAGCGUGAAAUGACUUUCACCUGCCCCCAGG-ACC---------------CA ...((.----(((...((((............((((((..((((...))))...)))))).......((.((((((....))))))))))))...))-).)---------------). ( -26.60) >DroPer_CAF1 29947 110 + 1 UCCAGC----UCCAACGGGCUGUUUAAUUUACGCGCUGCUAUUAUUUUAAUCUUUUGCGCUACACCCAGCGUGAAAUGACUUUCACCUGGCCCCG-GCGCCGGCCCCC-UGCUAUG-- ...(((----......((((((..........((...)).................(((((....((((.((((((....))))))))))....)-))))))))))..-.)))...-- ( -31.10) >consensus UCCAGC____UCCAACGGGCUGUUUAAUUUACGCGCUGCUAUUAUUUUAAUCUUUUGCGCUACACACAGCGUGAAAUGACUUUCACCUGGCCCCG_GCACCGGCUCC___________ ................((((............((((....................))))......(((.((((((....)))))))))))))......................... (-17.77 = -18.10 + 0.33)


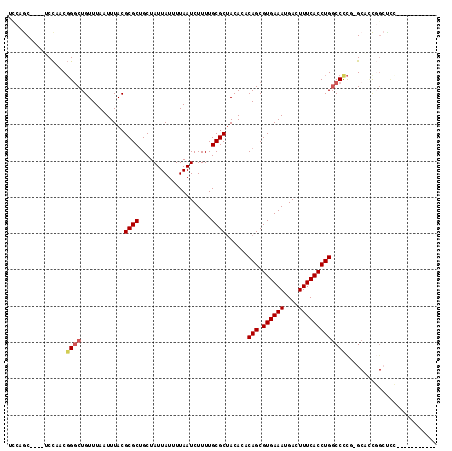
| Location | 8,200,403 – 8,200,508 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 78.41 |
| Mean single sequence MFE | -31.25 |
| Consensus MFE | -17.28 |
| Energy contribution | -17.27 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.55 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.533374 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 8200403 105 + 23771897 CUAUUAUUUUAAUCUUUUGCGCUACACACAGCGUGAAAUGACUUUCACCUGGCCUCGAGCAGCGGCUUCAAAGAUAUAGAAGCUAUAGC------UCCAAGUCGGGGUAGC ((((........(((((..((((......))))..))).))......((((((...((((.(..(((((.........)))))..).))------))...)))))))))). ( -32.30) >DroPse_CAF1 28088 106 + 1 CUAUUAUUUUAAUCUUUUGCGCUACACCCAGCGUGAAAUGACUUUCACCUGGCCCCG-GCGCCGGCCCCC-UGCUAUG---GUUACAACUACGGUUAAGGCUCGGGGUAGA ..................(((((....((((.((((((....))))))))))....)-))))..(((((.-.(((..(---....)(((....)))..)))..)))))... ( -33.90) >DroEre_CAF1 31353 111 + 1 CUAUUAUUUUAAUCUUUUGCGCUACACACAGCGUGAAAUGACUUUCACCUGGCCCCGAGCACCGACUUCAAAGAUAUAGAAGCUAUAGCUUCGGCUCCAAGUUGGGUUGGC ............(((((..((((......))))..))).))...(((((..((...((((...(....).........(((((....))))).))))...))..)).))). ( -29.30) >DroYak_CAF1 28785 111 + 1 CUAUUAUUUUAAUCUUUUGCGCUACACACAGCGUGAAAUGACUUUCACCUGGCCCCGAGCACCGGCUUCAAAGAUAUAGAAGCUAUAGCUUCGACUCCAAGUUGGGGUGGC ....................(((.....(((.((((((....)))))))))(((((((((...((((((.........))))))...)))).(((.....))))))))))) ( -30.50) >DroAna_CAF1 26835 95 + 1 CUAUUAUUUUAAUCUUUUGCGCUACACACAGCGUGAAAUGACUUUCACCUGCCCCCAGG-ACC---------------CAAGCUGUAGCUCCAGCUUCAAGUUGGGGUGGC ............(((((..((((......))))..))).)).......(..((((...(-((.---------------.((((((......))))))...)))))))..). ( -27.50) >DroPer_CAF1 29981 106 + 1 CUAUUAUUUUAAUCUUUUGCGCUACACCCAGCGUGAAAUGACUUUCACCUGGCCCCG-GCGCCGGCCCCC-UGCUAUG---GUUACAACUACGGUGAAGGCUCGGGGUAGA ..................(((((....((((.((((((....))))))))))....)-))))..(((((.-.(((.((---....))((....))...)))..)))))... ( -34.00) >consensus CUAUUAUUUUAAUCUUUUGCGCUACACACAGCGUGAAAUGACUUUCACCUGGCCCCGAGCACCGGCUUCA_AGAUAUAGAAGCUAUAGCUACGGCUCCAAGUCGGGGUAGC ............(((((..((((......))))..))).))..........(((((((((....))...............((....))............)))))))... (-17.28 = -17.27 + -0.01)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:43:49 2006