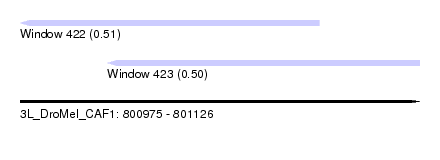

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 800,975 – 801,126 |
| Length | 151 |
| Max. P | 0.513924 |

| Location | 800,975 – 801,088 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.83 |
| Mean single sequence MFE | -28.96 |
| Consensus MFE | -21.98 |
| Energy contribution | -22.22 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.76 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.513924 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
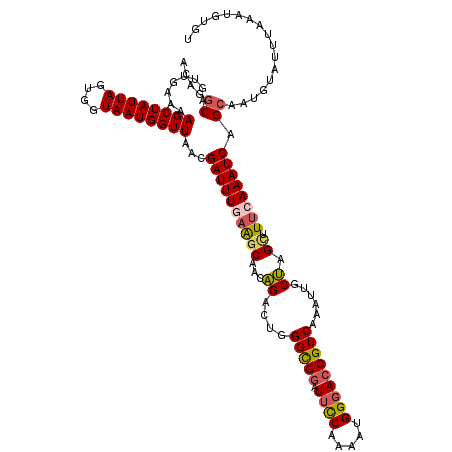
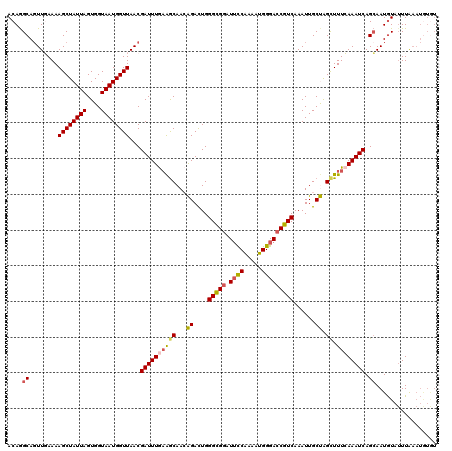
>3L_DroMel_CAF1 800975 113 - 23771897 ACAGGCACUUGAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGAAGCAACAGACUGGGCGGAUUUCAAAAUGGGACCGUCAAAUUGCUAGCUUUCAAAUCAGCAAUAUUAUUG------- ....((........((((((((....))))))))...((((((((((..(((....(((((.(..(.....)..))))))...)))...)).)))))))).))..........------- ( -27.30) >DroSec_CAF1 164527 120 - 1 ACAGGCAGUUGAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGAAGCAACAGACUGGGUGGAUUCCGAAAUGGGACCGUCAAAUUGCUGGCUUUCAAAUCAGCAAUGUAUUUAAAUGUGU ....(((.(((((.((((((((....))))))))...((((((((((..(((....(..((.((((.....))))))..)......))))).)))))))).........))))).))).. ( -28.40) >DroSim_CAF1 159181 120 - 1 ACAGGCAGUUGAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGAAGCAACAGACUGGGCGGAUUCCAUAAUGGGACCGUCAAAUUGCUGGCUUUCAAAUCAGCAAUGUAUUUAAAUGUAU ....(((.(((((.((((((((....))))))))...((((((((((..(((....(((((.((((.....)))))))))......))))).)))))))).........))))).))).. ( -33.10) >DroEre_CAF1 164124 120 - 1 ACAAGCAGUUUAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUAAGACAAGGGAUUGGGCGGAUUUCAAAGUGGGAACGUCAAAUUGCCAGCUUUAAAAUCAGAAAUGUAUUUAAAUGUGU ....(((((((...((((((((....))))))))..((((((........))))))((((..(..(.....)..).)))))))))))..((((((((..((....))..)))))).)).. ( -22.50) >DroYak_CAF1 164055 119 - 1 ACAGGCAGUUUAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGUAGCAAGGGAUUGGGCGGGU-CCAAAAUGGGACCGUCAAAUUGCCAGUUUUAAAAUCAGCAAUGUAUUUAAAUAUGU ...((((((((....(((((.(((.((........)).))).))))).........(((((..-((.....))..)))))))))))))...((((((..((....))..))))))..... ( -33.50) >consensus ACAGGCAGUUGAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGAAGCAACAGACUGGGCGGAUUCCAAAAUGGGACCGUCAAAUUGCUAGCUUUCAAAUCAGCAAUGUAUUUAAAUGUGU ....((........((((((((....))))))))...((((((((((...((....(((((.((((.....)))))))))......)).)).)))))))).))................. (-21.98 = -22.22 + 0.24)

| Location | 801,008 – 801,126 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 93.05 |
| Mean single sequence MFE | -33.06 |
| Consensus MFE | -27.92 |
| Energy contribution | -28.00 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.23 |
| Structure conservation index | 0.84 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
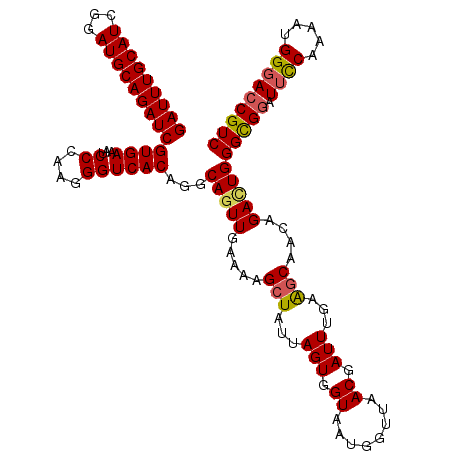
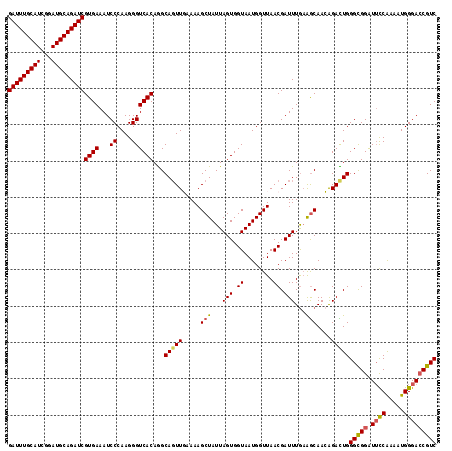
>3L_DroMel_CAF1 801008 118 - 23771897 GAUUUGCAUCGGAUGCAGAUCGUGAAAUCCCAAGGGUCACAGGCACUUGAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGAAGCAACAGACUGGGCGGAUUUCAAAAUGGGACCGUC (((((((((...)))))))))((.(((((.((((.((.....)).))))...((((((((....))))))))...)))))...)).........(((((.(..(.....)..)))))) ( -30.00) >DroSec_CAF1 164567 118 - 1 GAUUUGCAUCGGAUGCAGAUCGUGAAAUCCCAAGGGUCACAGGCAGUUGAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGAAGCAACAGACUGGGUGGAUUCCGAAAUGGGACCGUC (((((((((...)))))))))......(((((.(((((.((..(((((.....(((...(((.((........)).)))...)))....)))))..)))).)))....)))))..... ( -31.50) >DroSim_CAF1 159221 118 - 1 GAUUUGCAUCGGAUGCAGAUCGUGAAAUCCCAAGGGUCACAGGCAGUUGAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGAAGCAACAGACUGGGCGGAUUCCAUAAUGGGACCGUC (((((((((...)))))))))((((...((....))))))...(((((.....(((...(((.((........)).)))...)))....)))))(((((.((((.....))))))))) ( -35.20) >DroEre_CAF1 164164 118 - 1 GAUUUGCAUCGGAUGCAGAUCGUGAAAUCCCAAGGGUCACAAGCAGUUUAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUAAGACAAGGGAUUGGGCGGAUUUCAAAGUGGGAACGUC (((((((((...)))))))))....((((((....(((..(((..(((....((((((((....)))))))))))..)))..)))..)))))).((((..(..(.....)..).)))) ( -29.40) >DroYak_CAF1 164095 117 - 1 GAUUUGCAUCGGAUGCAGAUCGUGAAAUCCCAAGGGUCACAGGCAGUUUAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGUAGCAAGGGAUUGGGCGGGU-CCAAAAUGGGACCGUC (((((((((...)))))))))((..((((((....(..(((((..(((....((((((((....)))))))))))..)))))..)..))))))..)).(((-((......)))))... ( -39.20) >consensus GAUUUGCAUCGGAUGCAGAUCGUGAAAUCCCAAGGGUCACAGGCAGUUGAAAAGCUAUUAGUGGUAAUGGUUAACGAUUUGAAGCAACAGACUGGGCGGAUUCCAAAAUGGGACCGUC (((((((((...)))))))))((((...((....))))))...(((((.....(((...(((.((........)).)))...)))....)))))(((((.((((.....))))))))) (-27.92 = -28.00 + 0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:38:15 2006