
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,958,054 – 7,958,160 |
| Length | 106 |
| Max. P | 0.883013 |

| Location | 7,958,054 – 7,958,160 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 87.57 |
| Mean single sequence MFE | -32.25 |
| Consensus MFE | -23.59 |
| Energy contribution | -26.40 |
| Covariance contribution | 2.81 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.883013 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7958054 106 + 23771897 AGCCUCUUGGCAAGGUGUCUUUGGUCUAGGCUCACUCAGUCACUAACUCACUAGCAACUGAUUCGGCUUAGCCAGU---UCAGUUCCUGUUAUUUUUAGGCUGUGAACC---------- .(((....)))..........(((.(((((((.(.(((((..(((......)))..))))).).))))))))))((---(((...((((.......))))...))))).---------- ( -32.10) >DroSec_CAF1 3547 109 + 1 AGCCUCUUGGCAAGGUGUCUUUGGUCUAGGCUCACUCAGUCACUAACUCACCAGCAACUGAUUCGGCUUAGCCAGUUGUUCAGUUCCUGUUAUUUUUAGGCUCUGAACC---------- .(((....))).........((((.(((((((.(.(((((..((........))..))))).).)))))))))))..((((((..((((.......))))..)))))).---------- ( -35.10) >DroSim_CAF1 3548 109 + 1 AGCCUCUUGGCAAGGUGUCUUUGGUCUAGGCUCACUCAGUCACUAACUCACCAGCAACUGAUUCGGCUUAGCCAGUUGUUCAGUUCCUGUUAUUUUUAGGCUCUGAACC---------- .(((....))).........((((.(((((((.(.(((((..((........))..))))).).)))))))))))..((((((..((((.......))))..)))))).---------- ( -35.10) >DroYak_CAF1 3905 114 + 1 --CCUCUUGGCAAGGUGUCUUUGGUCUAGGCUCACUCAGUCACUAACUCACCAACCACUGAUUCGGCUCGGC---UUAGCCAGUUCCUGUUAUUUUUAGGCUCUGUAGCAGCAACAUCA --.....((((..((((...(((((...((((.....)))))))))..)))).....((((......)))).---...))))(((.(((((((...........))))))).))).... ( -26.70) >consensus AGCCUCUUGGCAAGGUGUCUUUGGUCUAGGCUCACUCAGUCACUAACUCACCAGCAACUGAUUCGGCUUAGCCAGUUGUUCAGUUCCUGUUAUUUUUAGGCUCUGAACC__________ .(((....))).........((((.(((((((.(.(((((..((........))..))))).).)))))))))))..((((((..((((.......))))..))))))........... (-23.59 = -26.40 + 2.81)

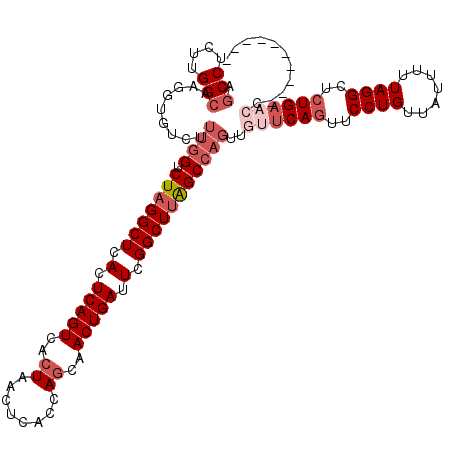

| Location | 7,958,054 – 7,958,160 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 87.57 |
| Mean single sequence MFE | -31.99 |
| Consensus MFE | -23.46 |
| Energy contribution | -25.15 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.672606 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7958054 106 - 23771897 ----------GGUUCACAGCCUAAAAAUAACAGGAACUGA---ACUGGCUAAGCCGAAUCAGUUGCUAGUGAGUUAGUGACUGAGUGAGCCUAGACCAAAGACACCUUGCCAAGAGGCU ----------(((((((((((...................---...))))........(((((..((((....))))..)))))))))))).................(((....))). ( -31.45) >DroSec_CAF1 3547 109 - 1 ----------GGUUCAGAGCCUAAAAAUAACAGGAACUGAACAACUGGCUAAGCCGAAUCAGUUGCUGGUGAGUUAGUGACUGAGUGAGCCUAGACCAAAGACACCUUGCCAAGAGGCU ----------.((((((..(((.........)))..))))))...((((((.((....(((((..((((....))))..)))))....)).))).)))..........(((....))). ( -34.70) >DroSim_CAF1 3548 109 - 1 ----------GGUUCAGAGCCUAAAAAUAACAGGAACUGAACAACUGGCUAAGCCGAAUCAGUUGCUGGUGAGUUAGUGACUGAGUGAGCCUAGACCAAAGACACCUUGCCAAGAGGCU ----------.((((((..(((.........)))..))))))...((((((.((....(((((..((((....))))..)))))....)).))).)))..........(((....))). ( -34.70) >DroYak_CAF1 3905 114 - 1 UGAUGUUGCUGCUACAGAGCCUAAAAAUAACAGGAACUGGCUAA---GCCGAGCCGAAUCAGUGGUUGGUGAGUUAGUGACUGAGUGAGCCUAGACCAAAGACACCUUGCCAAGAGG-- .....(..(((((....)))..........(((..(((((((..---(((.(((((......)))))))).)))))))..)))))..).(((......(((....)))......)))-- ( -27.10) >consensus __________GGUUCAGAGCCUAAAAAUAACAGGAACUGAACAACUGGCUAAGCCGAAUCAGUUGCUGGUGAGUUAGUGACUGAGUGAGCCUAGACCAAAGACACCUUGCCAAGAGGCU ...........((((((..(((.........)))..))))))...((((((.((....(((((..((((....))))..)))))....)).))).)))......((((.....)))).. (-23.46 = -25.15 + 1.69)


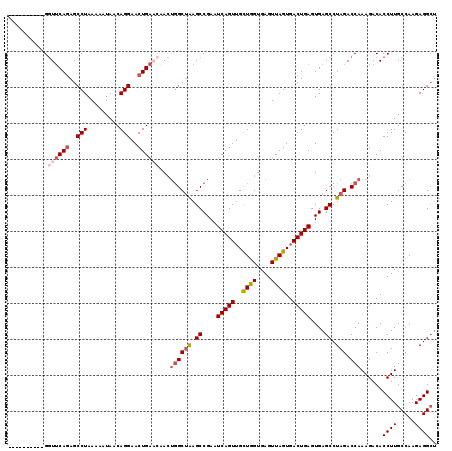
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:41:49 2006