
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,800,050 – 7,800,140 |
| Length | 90 |
| Max. P | 0.968223 |

| Location | 7,800,050 – 7,800,140 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 78.49 |
| Mean single sequence MFE | -37.68 |
| Consensus MFE | -29.00 |
| Energy contribution | -28.08 |
| Covariance contribution | -0.91 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.968223 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
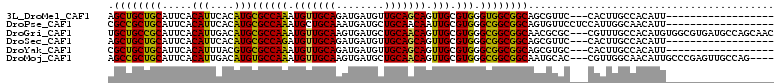
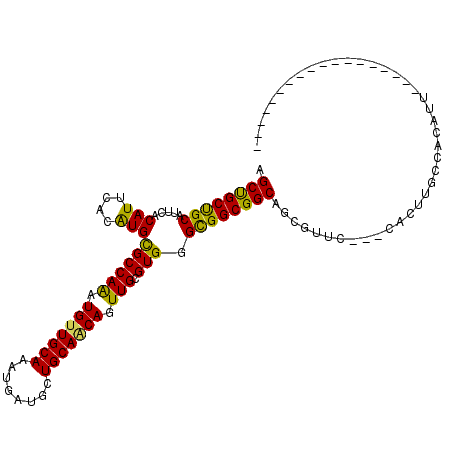
>3L_DroMel_CAF1 7800050 90 + 23771897 AGCUGCUGCAUUCACAUUCACAUGCGCCAAAUGUUGCAGAUGAUGUUGCAGCAGUUGCGUGGGUGGCGGCAGCGUUC---CACUUGCCACAUU------------------ ((((((((((...((((.((..((((.(....).))))..))))))))))))))))(.(..(((((((....))..)---))))..)).....------------------ ( -33.40) >DroPse_CAF1 10466 93 + 1 CGCCGCUGCAUUCACAUUCACAUGCGCCAAAUGCUGCAAAUGAUGCUGCAACAAUUGCGUGGGCGGCGGCAGUGUUCCUCCAUUGGCAACAUU------------------ .((((((((...........((((((.....(((.(((.....))).))).....)))))).))))))))((((((((......)).))))))------------------ ( -34.50) >DroGri_CAF1 26151 108 + 1 UGCUGCCGCAUUCACAUUGACAUGCGCCAAAUGUUGCAAGUGAUGCUGCAACAGUUGCGUGGGCGGCGGCAACGCGC---CGUUUGCCACAUGUGGCGUGAUGCCAGCAAC (((((((((..((.....))((((((.....(((((((........)))))))..)))))).)))))((((.(((((---((..((...))..))))))).)))))))).. ( -48.50) >DroSec_CAF1 9742 90 + 1 AGCUGCUGCAUUCACAUUCACAUGCGCCAGAUGUUGCAGAUGAUGUUGCAGCAGUUGCGUGGGCGGCGGCAGCGUUC---CACUUGCCACAUU------------------ .((((((((.....(((....)))((((.(.((((((((......))))))))....)...))))))))))))....---.............------------------ ( -32.10) >DroYak_CAF1 10308 90 + 1 CGCUGCUGCAUUCACAUUUACGUGCGCCAAAUGUUGCAGAUGAUGUUGCAGCAGUUGCGUGGGCGGCGGCAGCGUGC---CACUUGCCACAUU------------------ .((((((((...........(((.((((((.((((((((......)))))))).))).))).)))))))))))(((.---(....).)))...------------------ ( -35.90) >DroMoj_CAF1 27182 104 + 1 AGCCGCUGCAUUCACAUUGACAUGUGCCAAAUGUUGCAAGUGAUGCUGCAACAGUUGCGUGGGCGGCGGCAAUGCAC---CGUUGGCAACAUUGCCCGAGUUGCCAG---- .((((((((...(((((....)))))(((..(((((((........)))))))......))))))))))).......---...(((((((.((....))))))))).---- ( -41.70) >consensus AGCUGCUGCAUUCACAUUCACAUGCGCCAAAUGUUGCAAAUGAUGCUGCAACAGUUGCGUGGGCGGCGGCAGCGUUC___CACUUGCCACAUU__________________ .((((((((.....(((....)))((((((.(((((((........))))))).))).))).))))))))......................................... (-29.00 = -28.08 + -0.91)

| Location | 7,800,050 – 7,800,140 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 78.49 |
| Mean single sequence MFE | -35.58 |
| Consensus MFE | -23.31 |
| Energy contribution | -22.78 |
| Covariance contribution | -0.52 |
| Combinations/Pair | 1.33 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952332 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
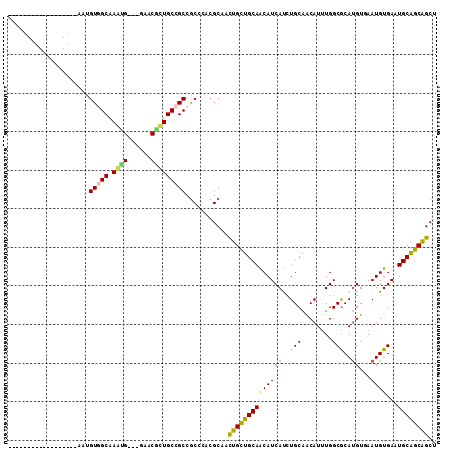
>3L_DroMel_CAF1 7800050 90 - 23771897 ------------------AAUGUGGCAAGUG---GAACGCUGCCGCCACCCACGCAACUGCUGCAACAUCAUCUGCAACAUUUGGCGCAUGUGAAUGUGAAUGCAGCAGCU ------------------...((((((.(((---...)))))))))...........(((((((((((((((.(((..(....)..))).))).))))...)))))))).. ( -31.40) >DroPse_CAF1 10466 93 - 1 ------------------AAUGUUGCCAAUGGAGGAACACUGCCGCCGCCCACGCAAUUGUUGCAGCAUCAUUUGCAGCAUUUGGCGCAUGUGAAUGUGAAUGCAGCGGCG ------------------..((((.((......))))))..(((((.((.....(((.((((((((......)))))))).))).(((((....)))))...)).))))). ( -32.60) >DroGri_CAF1 26151 108 - 1 GUUGCUGGCAUCACGCCACAUGUGGCAAACG---GCGCGUUGCCGCCGCCCACGCAACUGUUGCAGCAUCACUUGCAACAUUUGGCGCAUGUCAAUGUGAAUGCGGCAGCA (((((((.(((((((..(((((((.((((((---(((......)))))..........((((((((......)))))))))))).)))))))...)))).)))))))))). ( -46.40) >DroSec_CAF1 9742 90 - 1 ------------------AAUGUGGCAAGUG---GAACGCUGCCGCCGCCCACGCAACUGCUGCAACAUCAUCUGCAACAUCUGGCGCAUGUGAAUGUGAAUGCAGCAGCU ------------------...((((((.(((---...))))))))).((....))..(((((((((((((((.(((..(....)..))).))).))))...)))))))).. ( -32.10) >DroYak_CAF1 10308 90 - 1 ------------------AAUGUGGCAAGUG---GCACGCUGCCGCCGCCCACGCAACUGCUGCAACAUCAUCUGCAACAUUUGGCGCACGUAAAUGUGAAUGCAGCAGCG ------------------...((((...(((---((.....)))))...))))....((((((((((((...((((..(....)..))).)...))))...)))))))).. ( -31.40) >DroMoj_CAF1 27182 104 - 1 ----CUGGCAACUCGGGCAAUGUUGCCAACG---GUGCAUUGCCGCCGCCCACGCAACUGUUGCAGCAUCACUUGCAACAUUUGGCACAUGUCAAUGUGAAUGCAGCGGCU ----..(....)...((((((((.(((...)---))))))))))(((((.....(((.((((((((......)))))))).))).(((((....)))))......))))). ( -39.60) >consensus __________________AAUGUGGCAAAUG___GAACGCUGCCGCCGCCCACGCAACUGCUGCAACAUCAUCUGCAACAUUUGGCGCAUGUGAAUGUGAAUGCAGCAGCU .....................(((((.((((......)))))))))...........(((((((((((((((.(((........))).)))...))))...)))))))).. (-23.31 = -22.78 + -0.52)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:41:08 2006