
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,784,047 – 7,784,144 |
| Length | 97 |
| Max. P | 0.500000 |

| Location | 7,784,047 – 7,784,144 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 76.50 |
| Mean single sequence MFE | -30.92 |
| Consensus MFE | -16.87 |
| Energy contribution | -16.87 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.55 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7784047 97 + 23771897 GUGU--------UGGGGGUGCUAGUUUGCCAGUGGGCGUCAUUUAUCGUAUGAUGCGCAUUUCCGGCGCCACUCA---ACAGACUACAGCCAUAUAAACACC---AAGCAA ((((--------(..((.((.(((((((..((((((((((((.......)))))))((.......)).)))))..---.))))))))).)).....))))).---...... ( -33.30) >DroPse_CAF1 29074 90 + 1 G---------------U------GUGUUGUAGUGGGCGUCAUUUAUCGUAUGAUGCGCAUUUCCGGCGCCACUCAGCAACAGACGACAAGCAAACGGACACUCACAGACAG (---------------(------.((((((((((((((((((.......)))))))((.......)).)))))..)))))).))........................... ( -26.20) >DroSec_CAF1 24314 97 + 1 GUCU--------UGGGGGUGCUAGUUUGCCAGUGGGCGUCAUUUAUCGUAUGAUGCGCAUUUCCGGCGCCACUCA---ACAGACUACAGCCAUAUAAACACC---AAGCAA (.((--------(((.(((..(((((((..((((((((((((.......)))))))((.......)).)))))..---.)))))))..))).........))---)))).. ( -32.30) >DroEre_CAF1 24700 88 + 1 -----------------GUGCUAGUUUGGCAGUGGGCGUCAUUUAUCGUAUGAUGCGCAUUUCCGGCGCCACUCA---ACAGUCUACAGCCAUAUAAACACC---AAGCAA -----------------.((((.....((.((((.(((((((.......))))))).)))).))(((...(((..---..))).....)))...........---.)))). ( -25.00) >DroYak_CAF1 26327 105 + 1 GUGUGGGGGUGUGGGGGGUGCUAGUUUGCUAGUGGGCGUCAUUUAUCGUAUGAUGCGCAUUUCCGGCGCCACUCA---ACAGACUACAGCUACAUGAACAGC---AAGCAA ((.(((((((((.((..(((((((....)))....(((((((.......)))))))))))..)).))))).))))---))........(((........)))---...... ( -38.40) >DroPer_CAF1 29589 97 + 1 GUGU--------UUUGU------GUGUUGUAGUGGGCGUCAUUUAUCGUAUGAUGCGCAUUUCCGGCGCCACUCAGCAACAGACGACAAGCAAACGGACACUCACAGACAG ((((--------(((((------.((((((((((((((((((.......)))))))((.......)).)))))..)))))).))))..........))))).......... ( -30.30) >consensus GUGU________UGGGGGUGCUAGUUUGCCAGUGGGCGUCAUUUAUCGUAUGAUGCGCAUUUCCGGCGCCACUCA___ACAGACUACAGCCAUAUAAACACC___AAGCAA .......................((.....((((((((((((.......)))))))((.......)).))))).....))............................... (-16.87 = -16.87 + -0.00)

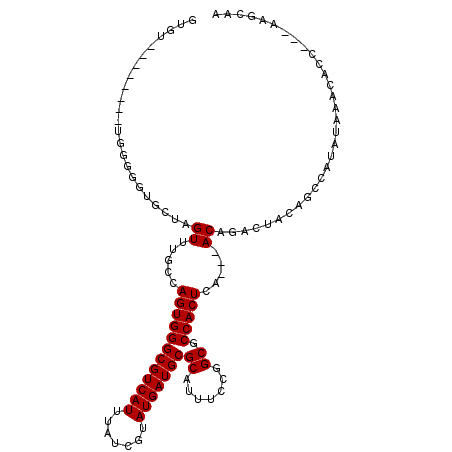

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:41:00 2006