
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,504,885 – 7,505,007 |
| Length | 122 |
| Max. P | 0.784898 |

| Location | 7,504,885 – 7,504,979 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 84.64 |
| Mean single sequence MFE | -23.16 |
| Consensus MFE | -14.40 |
| Energy contribution | -14.60 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.784898 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
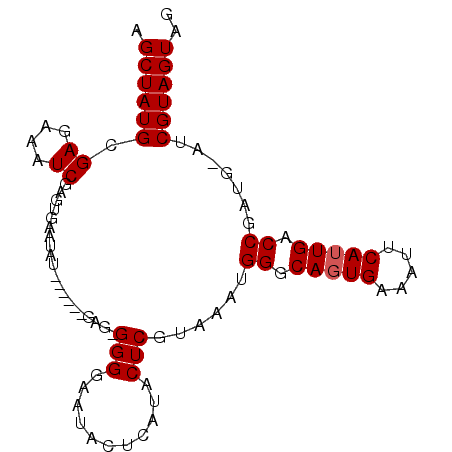
>3L_DroMel_CAF1 7504885 94 - 23771897 AGCUAUGCGAGAAAUCGAGUUAAUAUUGGUAUGAGGGGGGAAUACUCAUACUCGUAAAUGGGCAGUGAAAUUCAUUGACCGAUG-AUCGUAGUAA .(((((((((....)))......(((.((((((((.........)))))))).))).((.(((((((.....))))).)).)).-..)))))).. ( -27.40) >DroSec_CAF1 157627 87 - 1 AGCUAUGCGAGAAAUCGAGUGAAUAU------GAG-GGGGAAUACUCAUACUCGUAAAUGGGCAGUGAAAUUCAUUGACCGAUG-AUCGUAGUAG .(((((((((....)))...((.(((------(((-........)))))).))....((.(((((((.....))))).)).)).-..)))))).. ( -24.70) >DroSim_CAF1 149699 88 - 1 AGCUAUGCGAGAAAUCGAGUGAAUAU------GAG-GGGGAAUACUCAUACUCGUAAAUGGGCAGUGAAAUUCAUUGACCGAUGGAUCGUAGUAG .(((((((((....)))...((.(((------(((-........)))))).))((..((.(((((((.....))))).)).))..)))))))).. ( -24.80) >DroEre_CAF1 155381 87 - 1 AGCUAUGCGAGAAAUCAAGUGAAUAUAUGUAUGG--GGGGAAUG--AAUACUCGUAAAUGGGCAGUGAAAUUCAUUGACCG----AUCGUAGUAG .((((((.((....)).........(((.(((((--(.......--....)))))).)))(((((((.....))))).)).----..)))))).. ( -18.40) >DroYak_CAF1 151359 80 - 1 AGCUAUGCGAGAAAUCAAGUGAAUAUAU----GG--GGGGC-----AAUACUCGUAAGUGGUCACUGAAAUUCAUUGACCG----AUCGUAGUAG .((((((.((....)).........(((----((--(....-----....))))))..((((((.((.....)).))))))----..)))))).. ( -20.50) >consensus AGCUAUGCGAGAAAUCGAGUGAAUAU______GAG_GGGGAAUACUCAUACUCGUAAAUGGGCAGUGAAAUUCAUUGACCGAUG_AUCGUAGUAG .((((((.((....))....................(((...........)))......((.(((((.....))))).)).......)))))).. (-14.40 = -14.60 + 0.20)


| Location | 7,504,913 – 7,505,007 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 86.43 |
| Mean single sequence MFE | -20.47 |
| Consensus MFE | -14.75 |
| Energy contribution | -14.47 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.72 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.521343 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
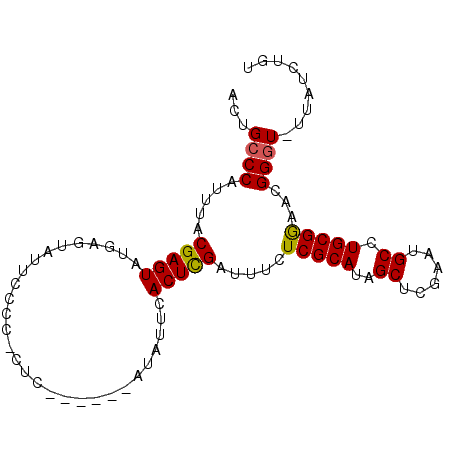
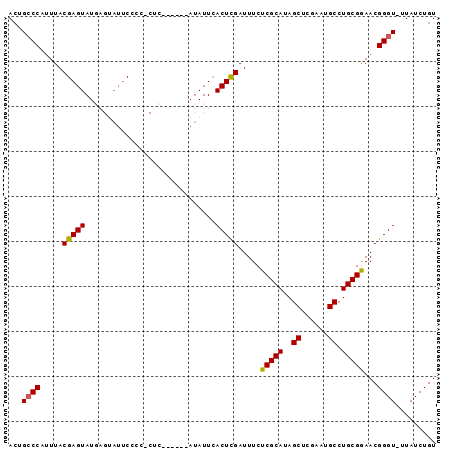
>3L_DroMel_CAF1 7504913 94 + 23771897 ACUGCCCAUUUACGAGUAUGAGUAUUCCCCCCUCAUACCAAUAUUAACUCGAUUUCUCGCAUAGCUCGAAUGCCUGCGGAACGGGU-UUAUCUGU ...((((......(.(((((((.........))))))))..............(((.((((..((......)).))))))).))))-........ ( -21.20) >DroSec_CAF1 157655 87 + 1 ACUGCCCAUUUACGAGUAUGAGUAUUCCCC-CUC------AUAUUCACUCGAUUUCUCGCAUAGCUCGAAUGCCUGCGGAACGGGU-UUAUCUGU ...((((......(((((((((........-)))------)))))).......(((.((((..((......)).))))))).))))-........ ( -24.70) >DroSim_CAF1 149728 87 + 1 ACUGCCCAUUUACGAGUAUGAGUAUUCCCC-CUC------AUAUUCACUCGAUUUCUCGCAUAGCUCGAAUGCCUGCGGAACGGGU-UUAUCUGU ...((((......(((((((((........-)))------)))))).......(((.((((..((......)).))))))).))))-........ ( -24.70) >DroEre_CAF1 155406 90 + 1 ACUGCCCAUUUACGAGUAUU--CAUUCCCC--CCAUACAUAUAUUCACUUGAUUUCUCGCAUAGCUCGAAUGCCUGCGAAACGGGU-UUAUCUGU ...((((......((((((.--........--........))))))..........(((((..((......)).)))))...))))-........ ( -16.23) >DroYak_CAF1 151384 84 + 1 AGUGACCACUUACGAGUAUU-----GCCCC--CC----AUAUAUUCACUUGAUUUCUCGCAUAGCUCGAAUGCCUGCGAAACGGGUUUUAUCUGU ...........(((..((..-----((((.--..----......((....))....(((((..((......)).)))))...))))..))..))) ( -15.50) >consensus ACUGCCCAUUUACGAGUAUGAGUAUUCCCC_CUC______AUAUUCACUCGAUUUCUCGCAUAGCUCGAAUGCCUGCGGAACGGGU_UUAUCUGU ...((((.....(((((.............................))))).....(((((..((......)).)))))...))))......... (-14.75 = -14.47 + -0.28)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:39:18 2006