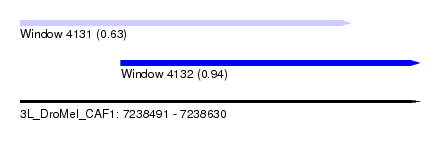
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,238,491 – 7,238,630 |
| Length | 139 |
| Max. P | 0.942837 |

| Location | 7,238,491 – 7,238,606 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 93.91 |
| Mean single sequence MFE | -45.40 |
| Consensus MFE | -39.95 |
| Energy contribution | -39.45 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.634954 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
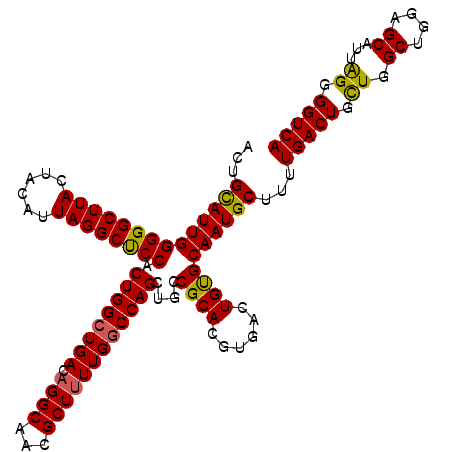
>3L_DroMel_CAF1 7238491 115 + 23771897 ACUGUAUUGGGGGCUUACUACAUUAGGCUCCACUGGCUGACAGGCACCGCUGUUGGCCAGCUGCCGCACGUGACUGUGCAAUGCUUUUGACUGCUGGCUGGAGCAUUGGGGGUCA .........((((((((......)))))))).(((((..((((......))))..)))))((.(((((((....)))))((((((((...(....)...)))))))))).))... ( -46.00) >DroSec_CAF1 19929 115 + 1 ACUGCAUUGGGGGCUUACUGCAUUAGGCUCCACUGGCUGACAGGCAACGCUUUUGGCCAGCUGCCGCACGUGACUGUGCAAUGCUUUUGACUGCUGGCUGGAGCAUUAGGGGUCA ...(((((.((((((((......)))))))).(((((..(.((((...)))))..))))).....(((((....))))))))))...(((((.(((((....))..))).))))) ( -42.40) >DroSim_CAF1 19981 115 + 1 AGUGCAUUGGGGGCUUAUUACAUUAGGCUCCACUGGAUGACAGGCAACGCUUUUGGCCAGCUGCCGCACGUGAUUGUGCAAUGCUUUUGACUGCUGGCUGGAGCAUUAGGGGUCA .((.((...((((((((......))))))))..)).))(((.(....)(((.(..((((((.((.(((((....)))))...))........))))))..)))).......))). ( -41.80) >DroEre_CAF1 20549 115 + 1 GGUGCAUUGGGGGCUUACUACAUUAGGCCCCACUGGCUGACAGGCACCGCUUUUGGCCAGCUGCCGCACGUGACUGCGCAAUGCUUUUGACUGUUGGCUGGAGCAUUGGGGGUCA .((((....((((((((......)))))))).(((((..(.((((...)))))..))))).....)))).(((((.(.(((((((((...(....)...)))))))))).))))) ( -51.40) >consensus ACUGCAUUGGGGGCUUACUACAUUAGGCUCCACUGGCUGACAGGCAACGCUUUUGGCCAGCUGCCGCACGUGACUGUGCAAUGCUUUUGACUGCUGGCUGGAGCAUUAGGGGUCA ...((((((((((((((......)))))))).((((((((.((((...))))))))))))....((((......))))))))))...(((((.((.((....))...)).))))) (-39.95 = -39.45 + -0.50)


| Location | 7,238,526 – 7,238,630 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 95.33 |
| Mean single sequence MFE | -43.84 |
| Consensus MFE | -37.08 |
| Energy contribution | -38.76 |
| Covariance contribution | 1.68 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.942837 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
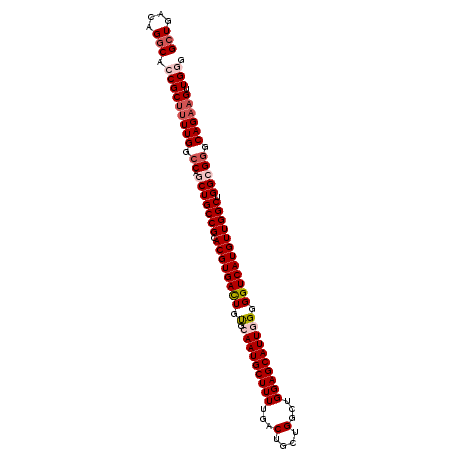
>3L_DroMel_CAF1 7238526 104 + 23771897 GCUGACAGGCACCGCUGUUGGCCAGCUGCCGCACGUGACUGUGCAAUGCUUUUGACUGCUGGCUGGAGCAUUGGGGGUCAUGUUGGCUGGCG-GCAGAAGUUGGG ((..((((......))))..))..((((((((((((((((.(.(((((((((...(....)...)))))))))).))))))))..)).))))-)).......... ( -45.20) >DroSec_CAF1 19964 105 + 1 GCUGACAGGCAACGCUUUUGGCCAGCUGCCGCACGUGACUGUGCAAUGCUUUUGACUGCUGGCUGGAGCAUUAGGGGUCAUGUUGGCUGGCGGGCAGAAGUUGGG (((....)))...(((((((.((.(((((((.((((((((.(..((((((((...(....)...)))))))).).)))))))))))).))))).))))))).... ( -43.50) >DroSim_CAF1 20016 105 + 1 GAUGACAGGCAACGCUUUUGGCCAGCUGCCGCACGUGAUUGUGCAAUGCUUUUGACUGCUGGCUGGAGCAUUAGGGGUCAUGUUGGCUGGCGGGCAGAAGUUGGG .......(....)(((((((.((.(((((((.((((((((.(..((((((((...(....)...)))))))).).)))))))))))).))))).))))))).... ( -39.90) >DroEre_CAF1 20584 105 + 1 GCUGACAGGCACCGCUUUUGGCCAGCUGCCGCACGUGACUGCGCAAUGCUUUUGACUGUUGGCUGGAGCAUUGGGGGUCAUGUUGGCUGGGGGGCAGAUGUUGGG .(..(((.((.((.((...(((((((....))((((((((.(.(((((((((...(....)...)))))))))).))))))))))))))).))))...)))..). ( -45.00) >DroYak_CAF1 21737 104 + 1 GCUGACAGGCACCGCUUUUGGCCAGCUGCCGCACGUGACUGUGCAAUGCUUUUGACUGUUGGCUGGAGCAUUGGGGGUCAUGUUGGCUGG-GGGCAGAAGUUGGG (((....))).(((((((((.((....((((.((((((((.(.(((((((((...(....)...)))))))))).))))))))))))...-)).)))))).))). ( -45.60) >consensus GCUGACAGGCACCGCUUUUGGCCAGCUGCCGCACGUGACUGUGCAAUGCUUUUGACUGCUGGCUGGAGCAUUGGGGGUCAUGUUGGCUGGCGGGCAGAAGUUGGG (((....))).(((((((((.((.(((((((.((((((((.(.(((((((((...(....)...)))))))))).)))))))))))).))))).)))))).))). (-37.08 = -38.76 + 1.68)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:38:07 2006