| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,229,103 – 7,229,201 |
| Length | 98 |
| Max. P | 0.986879 |

| Location | 7,229,103 – 7,229,201 |
|---|---|
| Length | 98 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 83.54 |
| Mean single sequence MFE | -28.42 |
| Consensus MFE | -25.87 |
| Energy contribution | -26.53 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.772589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7229103 98 + 23771897 GCAUAUUUUU--------------UUUGGGGGCGCGGUUUAGAAAGUUGUGCUCCCCCUCCAUGCCAUGGAUGUGGCCAAAAAAAAAAAAAAAAAACUGGGCCAUGGCAUUC ..........--------------...((((((((..((.....))..)))))))).....(((((((((......(((..................))).))))))))).. ( -28.77) >DroSec_CAF1 10802 105 + 1 GCAUAUUUUUCUUUUUUUUUUUUAUUUGGGGGCGCGGUUUAGAAAGUUUGGCUCCCCCUCCAUGCCAUGGAUGUGGCCGAAAA-------AAAAAACUGGGCCAUGGCAUUC ((.(((..(((.(((((((((((....(((((.((.(...........).)).))))).....(((((....))))).)))))-------))))))..)))..))))).... ( -27.60) >DroSim_CAF1 10721 103 + 1 GCAUAUUUUUCUUGUU-UUUUUCAUCUGGGGGCGCGGUUUAGAAAGUUUUGCUCCCCCUCCAUGCCAUGGAUGUGGCCGAAAA--------AAAAACUGGGCCAUGGCAUUC ((.(((..(((...((-((((((....(((((.((((...........)))).))))).....(((((....))))).)))))--------)))....)))..))))).... ( -28.90) >consensus GCAUAUUUUUCUU_UU_UUUUU_AUUUGGGGGCGCGGUUUAGAAAGUUUUGCUCCCCCUCCAUGCCAUGGAUGUGGCCGAAAA_______AAAAAACUGGGCCAUGGCAUUC ...........................(((((.((((...........)))).)))))...(((((((((..((.....................))....))))))))).. (-25.87 = -26.53 + 0.67)


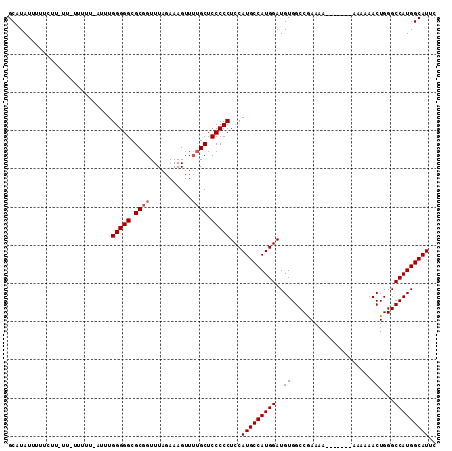
| Location | 7,229,103 – 7,229,201 |
|---|---|
| Length | 98 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 83.54 |
| Mean single sequence MFE | -27.16 |
| Consensus MFE | -25.82 |
| Energy contribution | -25.82 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.95 |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.986879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7229103 98 - 23771897 GAAUGCCAUGGCCCAGUUUUUUUUUUUUUUUUUUGGCCACAUCCAUGGCAUGGAGGGGGAGCACAACUUUCUAAACCGCGCCCCCAAA--------------AAAAAUAUGC ..(((((((((.((((................))))......)))))))))...(((((.((...............)).)))))...--------------.......... ( -27.55) >DroSec_CAF1 10802 105 - 1 GAAUGCCAUGGCCCAGUUUUUU-------UUUUCGGCCACAUCCAUGGCAUGGAGGGGGAGCCAAACUUUCUAAACCGCGCCCCCAAAUAAAAAAAAAAAAGAAAAAUAUGC ..(((((((((((.((......-------..)).)))))......))))))...(((((.((...............)).)))))........................... ( -26.76) >DroSim_CAF1 10721 103 - 1 GAAUGCCAUGGCCCAGUUUUU--------UUUUCGGCCACAUCCAUGGCAUGGAGGGGGAGCAAAACUUUCUAAACCGCGCCCCCAGAUGAAAAA-AACAAGAAAAAUAUGC ....((....))...((((((--------(((((.((((......)))).....(((((.((...............)).))))).)).))))))-)))............. ( -27.16) >consensus GAAUGCCAUGGCCCAGUUUUUU_______UUUUCGGCCACAUCCAUGGCAUGGAGGGGGAGCAAAACUUUCUAAACCGCGCCCCCAAAU_AAAAA_AA_AAGAAAAAUAUGC ..(((((((((((.((...............)).)))))......))))))...(((((.((...............)).)))))........................... (-25.82 = -25.82 + 0.00)


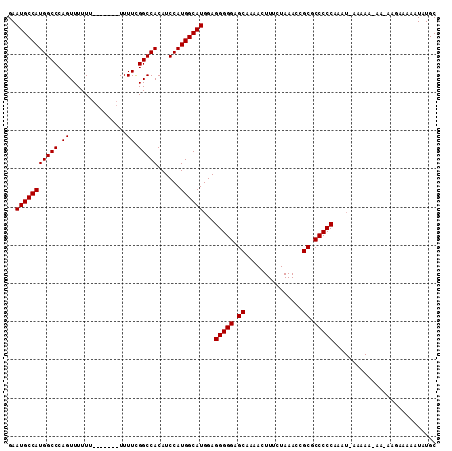
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:37:32 2006