
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,221,590 – 7,221,680 |
| Length | 90 |
| Max. P | 0.925047 |

| Location | 7,221,590 – 7,221,680 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 76.72 |
| Mean single sequence MFE | -39.47 |
| Consensus MFE | -20.67 |
| Energy contribution | -21.83 |
| Covariance contribution | 1.17 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.52 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925047 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7221590 90 + 23771897 GUAGA---------GCUGCUGGUUCUGCUGAUACUGGCCGGCCGGGUCCUGCGGAGC---UCCCAUAGGUGUGUCCUCCAGCAUCGCCAGCAUGCUGGAGUU (((((---------((.....))))))).(((((..(((....(((..((....)).---.)))...))))))))((((((((((....).))))))))).. ( -35.50) >DroPse_CAF1 3958 96 + 1 GUACAAUUGACUUUGCUGCUGCUGCGGCUGCGGCUGGC---CCGUCUCUUGGGGACA---GCCCAUGGGCGUAUCCUCCAGCAUCGCCAGCAUGCUGGAGUU ((((..........(((((.((....)).)))))..((---((((...(((....))---)...)))))))))).((((((((((....).))))))))).. ( -40.90) >DroEre_CAF1 3255 90 + 1 GUAGA---------GCUGCUGGUUCUGCUGAUGCUGACCGGCUGUGUCCUGCGGAUA---UCCCAUAGGUGUGUCCUCCAGCAUCGCCAGCAUGCUGGAGUU ((((.---------.(.(((((((..((....)).)))))))...)..))))(((((---..((...))..)))))(((((((((....).))))))))... ( -39.20) >DroYak_CAF1 3255 90 + 1 GUAUA---------GCUGCUGGUGCUGCUGCUGUUGACCGGCCGUGUCCUGCGGAUA---ACCCAUAGGUGUAUCCUCCAGCAUCGCCAGCAUGCUGGAGUU ...((---------((((((((((.(((((..((.((((....).)))..))(((((---(((....))).)))))..))))).)))))))).))))..... ( -37.20) >DroAna_CAF1 3168 87 + 1 GUAUA---------GCUGCUGGUGUUGCUG------CACGGACGAAUCGUGGGUGUAGCCCCCCAUCGGCGUGUCCUCCAGCAUCGCCAGCAUGCUGGAGUU ...((---------((((((((((.(((((------...(((((..(((((((........)))).)))..)))))..))))).)))))))).))))..... ( -43.10) >DroPer_CAF1 3784 96 + 1 GUACAAUUGACUUUGCUGCUGCUGCGGCUGCGGCUGGC---CCGUCUCUUGGGGACA---GCCCAUGGGCGUAUCCUCCAGCAUCGCCAGCAUGCUGGAGUU ((((..........(((((.((....)).)))))..((---((((...(((....))---)...)))))))))).((((((((((....).))))))))).. ( -40.90) >consensus GUACA_________GCUGCUGGUGCUGCUGCUGCUGGCCGGCCGUGUCCUGCGGAUA___GCCCAUAGGCGUAUCCUCCAGCAUCGCCAGCAUGCUGGAGUU ..............((((((((((.(((((......................((........))..(((.....))).))))).)))))))).))....... (-20.67 = -21.83 + 1.17)

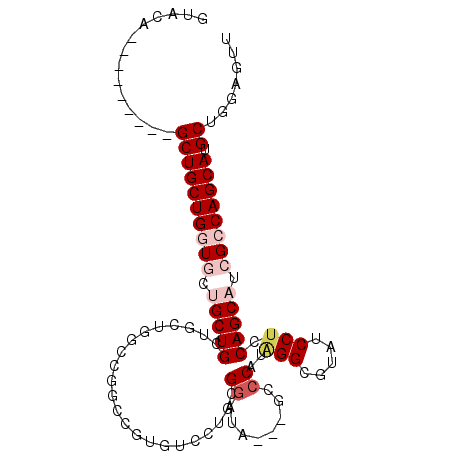

| Location | 7,221,590 – 7,221,680 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 76.72 |
| Mean single sequence MFE | -34.08 |
| Consensus MFE | -18.45 |
| Energy contribution | -19.15 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.686613 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7221590 90 - 23771897 AACUCCAGCAUGCUGGCGAUGCUGGAGGACACACCUAUGGGA---GCUCCGCAGGACCCGGCCGGCCAGUAUCAGCAGAACCAGCAGC---------UCUAC ......(((.((((((.((((((((.((.....(((.(((..---...))).)))......))..)))))))).......))))))))---------).... ( -30.80) >DroPse_CAF1 3958 96 - 1 AACUCCAGCAUGCUGGCGAUGCUGGAGGAUACGCCCAUGGGC---UGUCCCCAAGAGACGG---GCCAGCCGCAGCCGCAGCAGCAGCAAAGUCAAUUGUAC ..((((((((((....).)))))))))(((..((...(((((---((.(((........))---).))))).))...)).((....))...)))........ ( -35.90) >DroEre_CAF1 3255 90 - 1 AACUCCAGCAUGCUGGCGAUGCUGGAGGACACACCUAUGGGA---UAUCCGCAGGACACAGCCGGUCAGCAUCAGCAGAACCAGCAGC---------UCUAC ......(((.((((((.((((((((.((.....(((.(((..---...))).)))......))..)))))))).......))))))))---------).... ( -30.20) >DroYak_CAF1 3255 90 - 1 AACUCCAGCAUGCUGGCGAUGCUGGAGGAUACACCUAUGGGU---UAUCCGCAGGACACGGCCGGUCAACAGCAGCAGCACCAGCAGC---------UAUAC ......(((.((((((.(.(((((..(((((.(((....)))---)))))....(((.(....)))).....))))).).))))))))---------).... ( -34.20) >DroAna_CAF1 3168 87 - 1 AACUCCAGCAUGCUGGCGAUGCUGGAGGACACGCCGAUGGGGGGCUACACCCACGAUUCGUCCGUG------CAGCAACACCAGCAGC---------UAUAC ......(((.((((((.(.(((((.(((((....((.((((........))))))....)))).).------))))).).))))))))---------).... ( -37.50) >DroPer_CAF1 3784 96 - 1 AACUCCAGCAUGCUGGCGAUGCUGGAGGAUACGCCCAUGGGC---UGUCCCCAAGAGACGG---GCCAGCCGCAGCCGCAGCAGCAGCAAAGUCAAUUGUAC ..((((((((((....).)))))))))(((..((...(((((---((.(((........))---).))))).))...)).((....))...)))........ ( -35.90) >consensus AACUCCAGCAUGCUGGCGAUGCUGGAGGACACACCUAUGGGA___UAUCCCCAGGACACGGCCGGCCAGCAGCAGCAGCACCAGCAGC_________UAUAC .......((.((((((...(((((..((.....))...(((......)))......................)))))...)))))))).............. (-18.45 = -19.15 + 0.70)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:37:18 2006