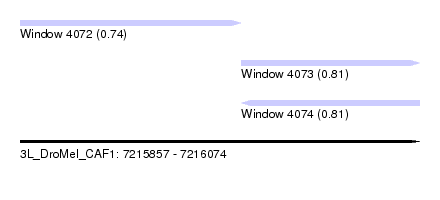

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,215,857 – 7,216,074 |
| Length | 217 |
| Max. P | 0.813329 |

| Location | 7,215,857 – 7,215,977 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -48.33 |
| Consensus MFE | -46.00 |
| Energy contribution | -45.90 |
| Covariance contribution | -0.10 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.744272 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
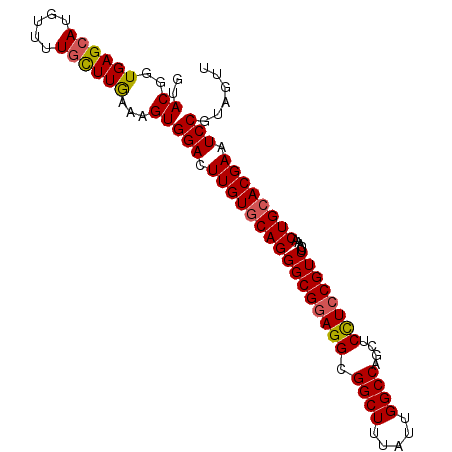
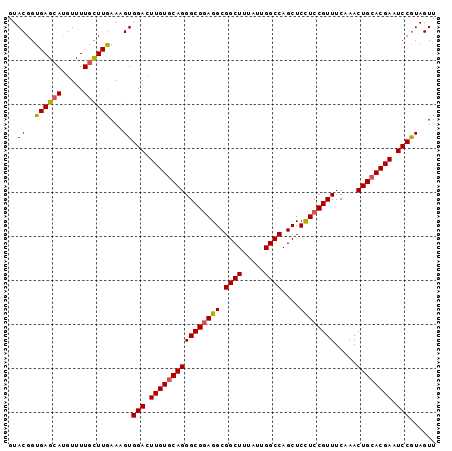
>3L_DroMel_CAF1 7215857 120 + 23771897 AAGGAUCCGGCCCUGGCAAAGAGUCGAAUCUUCUUGGGCAACCUGCCGGUUUGCACGCGCGAGGAGCUGGUGAGCAUUUGCCAGCCAUAUGGCAAAGUGCUCGGUUCGAUGGUCCAGAAG ..(((((..((....)).....((((((((.....((....)).((((((((((....))))...))))))((((((((((((......)))).)))))))))))))))))))))..... ( -49.70) >DroSim_CAF1 9367 120 + 1 AAGGACCCGGCCCUGGCCAAGAGUCGAAUCUUCUUGGGCAACCUGCCGGUUUGCACGCGCGAGGAGCUGGUGAGCAUUUGUCAGCCGUAUGGCAAAGUGCUUGGUUCGAUGGUCCAGAAG ..(((((.(((....)))....((((((((.....((....)).((((((((((....))))...))))))((((((((((((......)))).)))))))))))))))))))))..... ( -50.50) >DroYak_CAF1 5258 120 + 1 AAGGAUCCGGCGCUUGCCAAGAGUCGUAUCUUCUUGGGCAACCUACCGGUUUGCACGCGCGAGGAGCUGGUGAGCAUUUGCCAGCCGUAUGGCAAGGUCCUUGGUUCGAUGGUCCAGAAA ..(((((..((((...(((((((.......)))))))((((((....)).))))..))))...((((..(.(..(.(((((((......))))))))..))..))))...)))))..... ( -44.80) >consensus AAGGAUCCGGCCCUGGCCAAGAGUCGAAUCUUCUUGGGCAACCUGCCGGUUUGCACGCGCGAGGAGCUGGUGAGCAUUUGCCAGCCGUAUGGCAAAGUGCUUGGUUCGAUGGUCCAGAAG ..(((((.(((....)))....((((((((.....((....)).((((((((((....))))...))))))((((((((((((......))))))).))))))))))))))))))..... (-46.00 = -45.90 + -0.10)

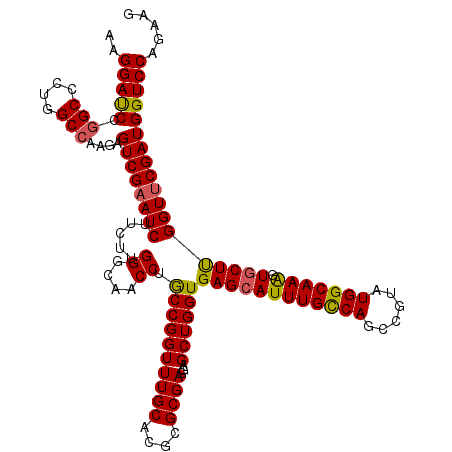
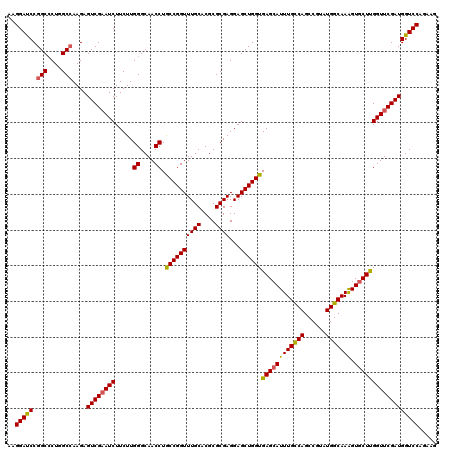
| Location | 7,215,977 – 7,216,074 |
|---|---|
| Length | 97 |
| Sequences | 3 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 93.47 |
| Mean single sequence MFE | -29.17 |
| Consensus MFE | -24.25 |
| Energy contribution | -24.70 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.808355 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7215977 97 + 23771897 AACUACGGAUUCGUGCAGUUUGAAACGGAGGAGCUGGCCAAUAAAGCCGCCUCCGCCCUGCACAAGUCCACUUUUAAGCAAAACAUGCUCACCGUAC ...((((((((.((((((........((.((((.((((.......)))).)))).)))))))).))))).......((((.....))))....))). ( -32.30) >DroSim_CAF1 9487 97 + 1 AACUAUGGAUUCGUGCAGUUUGAAACGGAGGAGCUGGCCAAUAAAGCCGCCUUCGCCCUGCACAAGUCCACCUUCAAGCAAAACAUGCUCACCGUAC .....((((((.((((((........((.((((.((((.......)))).)))).)))))))).))))))......((((.....))))........ ( -28.80) >DroYak_CAF1 5378 97 + 1 AACUACGGAUUCGUUCAGUUUGAAACGGAAGAGCUGGCCAAUAAAGCCGCCUCCGCCCUGCACAAGUCCACGUUCAAACAAAACGUCCUCACCGUAC ...(((((...((((..(((((((..(((.(((.((((.......)))).))).((...)).....)))...)))))))..))))......))))). ( -26.40) >consensus AACUACGGAUUCGUGCAGUUUGAAACGGAGGAGCUGGCCAAUAAAGCCGCCUCCGCCCUGCACAAGUCCACCUUCAAGCAAAACAUGCUCACCGUAC ......(((((.((((((........((.((((.((((.......)))).)))).)))))))).)))))............................ (-24.25 = -24.70 + 0.45)

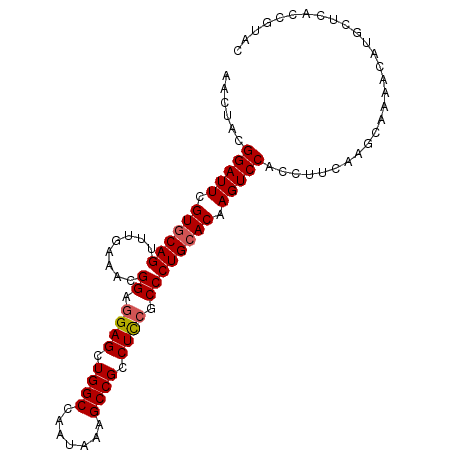
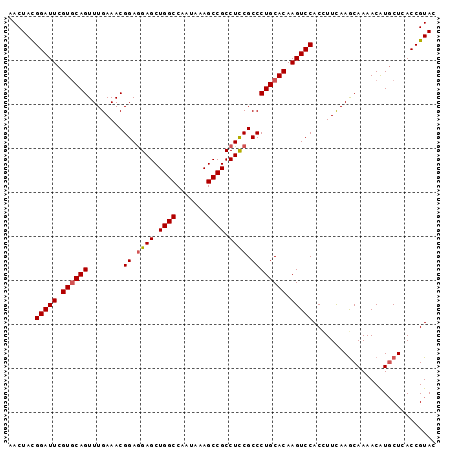
| Location | 7,215,977 – 7,216,074 |
|---|---|
| Length | 97 |
| Sequences | 3 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 93.47 |
| Mean single sequence MFE | -36.60 |
| Consensus MFE | -30.99 |
| Energy contribution | -31.33 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.813329 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7215977 97 - 23771897 GUACGGUGAGCAUGUUUUGCUUAAAAGUGGACUUGUGCAGGGCGGAGGCGGCUUUAUUGGCCAGCUCCUCCGUUUCAAACUGCACGAAUCCGUAGUU .(((..((((((.....)))))).....(((.((((((((((((((((.((((.....))))....)))))))).....)))))))).))))))... ( -40.30) >DroSim_CAF1 9487 97 - 1 GUACGGUGAGCAUGUUUUGCUUGAAGGUGGACUUGUGCAGGGCGAAGGCGGCUUUAUUGGCCAGCUCCUCCGUUUCAAACUGCACGAAUCCAUAGUU ...(..((((((.....))))))..)(((((.((((((((.((....))((((.....)))).................)))))))).))))).... ( -35.60) >DroYak_CAF1 5378 97 - 1 GUACGGUGAGGACGUUUUGUUUGAACGUGGACUUGUGCAGGGCGGAGGCGGCUUUAUUGGCCAGCUCUUCCGUUUCAAACUGAACGAAUCCGUAGUU .(((((......(((((.((((((((.((.(....).)).)((((((((((((.....)))).)))..)))))))))))).)))))...)))))... ( -33.90) >consensus GUACGGUGAGCAUGUUUUGCUUGAAAGUGGACUUGUGCAGGGCGGAGGCGGCUUUAUUGGCCAGCUCCUCCGUUUCAAACUGCACGAAUCCGUAGUU ..((..((((((.....))))))...))(((.((((((((((((((((.((((.....))))....)))))))).....)))))))).)))...... (-30.99 = -31.33 + 0.34)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:37:14 2006