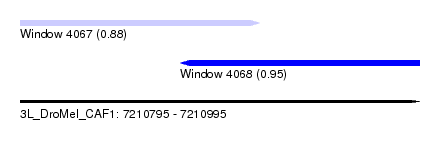
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,210,795 – 7,210,995 |
| Length | 200 |
| Max. P | 0.950666 |

| Location | 7,210,795 – 7,210,915 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.53 |
| Mean single sequence MFE | -44.25 |
| Consensus MFE | -38.39 |
| Energy contribution | -39.08 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.880685 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7210795 120 + 23771897 CCCAUUCCAUCCGCUUGGCACCGUCUGCGAGCCUAAUGUGCCACUUAAGACGCAUUUAUUGUCUUUCGUUGCUGCUGCCGGCGUUCUUGCCGCGCGGCGGCUGUUGGUGGUGCCGCAUUU ............((..((((((..(.((.((((......((.((..((((((.......))))))..)).)).((((((((((....))))).))))))))))).)..)))))))).... ( -50.00) >DroSec_CAF1 3927 120 + 1 CCCAUUCCAUCCGCUUGGCACCGUCUGCGAGCCUAAUGUGCCACUUAAGACGCAUUUAUUGUCUUUCGUUGCUGCUGCCGGCGUUCUUGCCGCACGGCGGCUGUUGUUGGUGCCGCAUUU ............((..(((((((...((.((((......((.((..((((((.......))))))..)).)).((((.(((((....)))))..))))))))))...))))))))).... ( -43.00) >DroSim_CAF1 3953 120 + 1 CCCAUUCCAUCCGCUUGGCACCGUCUGCGAGCCUAAUGUGCCACUUAAGACGCAUUUAUUGUCUUUCGUUCCUGCUGCCGGCGUUCUUGCCGCACGGCGGCUGUUGUUGGUGCCGCAUUU ............((..(((((((...((.((((...(((((.((..((((((.......))))))..))..........((((....)))))))))..))))))...))))))))).... ( -39.40) >DroEre_CAF1 500 120 + 1 CCCAUUCCAUCCGCUUGGCACCGUCUGCGAGCCUAAUGUGCCACUUAAGACGCAUUUAUUGUCUUUCGUUGUUGCUGCUGCUGUUCUUGCAGCACGGCGGCUGUUGGUGGUGCCGCAUUU ............((..((((((..(.((.((((......((.((..((((((.......))))))..)).)).((((.((((((....)))))))))))))))).)..)))))))).... ( -44.60) >consensus CCCAUUCCAUCCGCUUGGCACCGUCUGCGAGCCUAAUGUGCCACUUAAGACGCAUUUAUUGUCUUUCGUUGCUGCUGCCGGCGUUCUUGCCGCACGGCGGCUGUUGGUGGUGCCGCAUUU ............((..(((((((((.((.((((......((.((..((((((.......))))))..)).)).((((.(((((....)))))..)))))))))).))))))))))).... (-38.39 = -39.08 + 0.69)

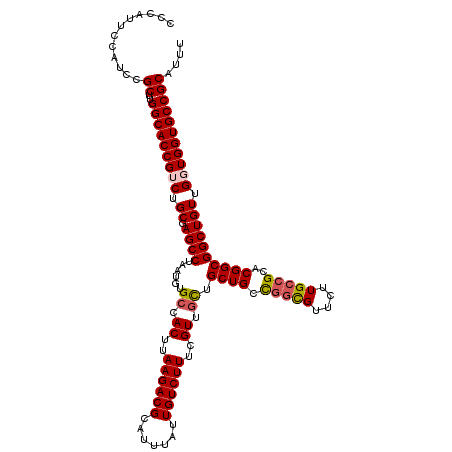
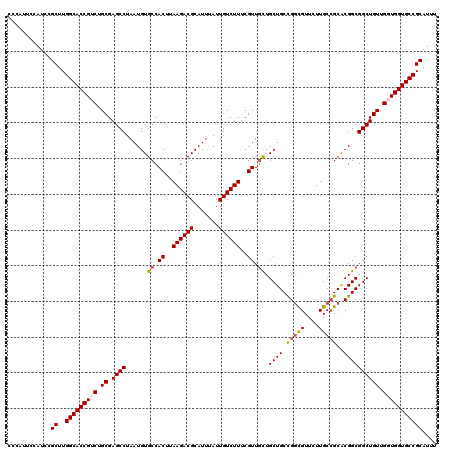
| Location | 7,210,875 – 7,210,995 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.36 |
| Mean single sequence MFE | -41.03 |
| Consensus MFE | -33.96 |
| Energy contribution | -35.52 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.950666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7210875 120 - 23771897 GCAAGUUGGGCUUGGAAAGCGGUGCAGCGAAAGCCAACACACGACCGCCAUUUUGUCAACAAUGGCGUCAAUUAAAUGGAAAAUGCGGCACCACCAACAGCCGCCGCGCGGCAAGAACGC ((..((((((((.....)))(((((.(((....(((......(((.((((((........))))))))).......)))....))).))))).))))).(((((...)))))......)) ( -44.42) >DroSec_CAF1 4007 120 - 1 GCAAGUUGGGCUUGGAAAGCGGUGCAGCGUAAGCCAACACACGACCGCCAUUUUGUCAACAAUGGCGUCAAUUAAAUGGAAAAUGCGGCACCAACAACAGCCGCCGUGCGGCAAGAACGC ((..((((.(((.....)))(((((.((((...(((......(((.((((((........))))))))).......)))...)))).)))))..)))).(((((...)))))......)) ( -42.82) >DroSim_CAF1 4033 120 - 1 GCAAGUUGGGCUUGGAAAGCGGUGCAGCGAAAGCCAACACACGACCGCCAUUUUGUCAACAAUGGCGUCAAUUAAAUGGAAAAUGCGGCACCAACAACAGCCGCCGUGCGGCAAGAACGC ((..((((.(((.....)))(((((.(((....(((......(((.((((((........))))))))).......)))....))).)))))..)))).(((((...)))))......)) ( -41.12) >DroEre_CAF1 580 112 - 1 GCAACUUGGGU--------GGGUGCAGCGAAAGCCAACACACAACCGCCAUUUUGUCAACAAUGGCGUCAAUUAAAUGGAAAAUGCGGCACCACCAACAGCCGCCGUGCUGCAAGAACAG ....(((((((--------((.(((.(((....(((.........(((((((........))))))).........)))....))).))))))))..((((......))))))))..... ( -35.77) >consensus GCAAGUUGGGCUUGGAAAGCGGUGCAGCGAAAGCCAACACACGACCGCCAUUUUGUCAACAAUGGCGUCAAUUAAAUGGAAAAUGCGGCACCAACAACAGCCGCCGUGCGGCAAGAACGC ....((((((((.....)))(((((.(((....(((.........(((((((........))))))).........)))....))).))))).))))).(((((...)))))........ (-33.96 = -35.52 + 1.56)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:37:08 2006