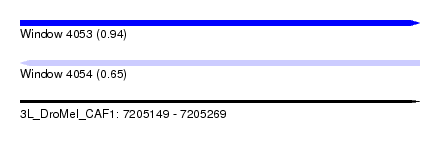

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 7,205,149 – 7,205,269 |
| Length | 120 |
| Max. P | 0.943629 |

| Location | 7,205,149 – 7,205,269 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.20 |
| Mean single sequence MFE | -31.57 |
| Consensus MFE | -26.30 |
| Energy contribution | -26.55 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.943629 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7205149 120 + 23771897 UGCCGCAGACAUAUUGUGUCUGCAAAACUUUGUUGACAACAAAGGAAAAAGGAAAACAGGAAGGCAAUGAGCAAACGCAAACAGAUGGGGAAAAGUGUCUCGAUUUUCAGUUACCGCCCC ((((((((((((...))))))))....((((((.....))))))..................))))....((....))........((((....(((.((.(.....))).)))..)))) ( -30.80) >DroSec_CAF1 18481 118 + 1 UGCCGCAGACAUAUUGUGUCUGCAAAACUUUGUUGACAACAAAGGAAAAAGGAAAACAGCAAGGCAAUGAGCAAACGCAAACAGAUGGGGAAAAGUGUCUCGAUUUUCAGUUACCU--CC ((((((((((((...))))))))....((((((.....))))))..................))))..(((..(((......((((.((((......)))).))))...)))..))--). ( -29.90) >DroEre_CAF1 18933 105 + 1 UGCCGCAGACAUAUUGUGUCUGCGAAACUUUGUUGACAACAAAGGGAAAAGGAAAACAGCAAGGCAAUGAGCA------CAGAGCUGGGGAAAAGUGUCUCGAUUUUCAGU--------- ...(((((((((...)))))))))...((((((.....)))))).(((((......((((...((.....)).------....))))((((......))))..)))))...--------- ( -29.90) >DroYak_CAF1 16087 114 + 1 UGCCGCAGACAUAUUGUGUCUGCGAAACUUUGUUGACAACAAAGCAAAAAGGAAAACAGCAGGGCAAUGAGCA------AACAGAUGGGGAAAAGUGUCUCGAUUUUCAGUUCUCGCCCC ((((((((((((...)))))))))...((((((.....))))))..............)))((((...((((.------...((((.((((......)))).))))...))))..)))). ( -35.70) >consensus UGCCGCAGACAUAUUGUGUCUGCAAAACUUUGUUGACAACAAAGGAAAAAGGAAAACAGCAAGGCAAUGAGCA______AACAGAUGGGGAAAAGUGUCUCGAUUUUCAGUUACCG__CC ((((((((((((...))))))))....((((((.....))))))..................))))................((((.((((......)))).)))).............. (-26.30 = -26.55 + 0.25)

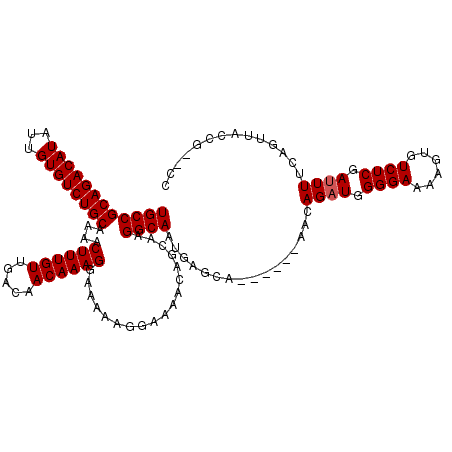
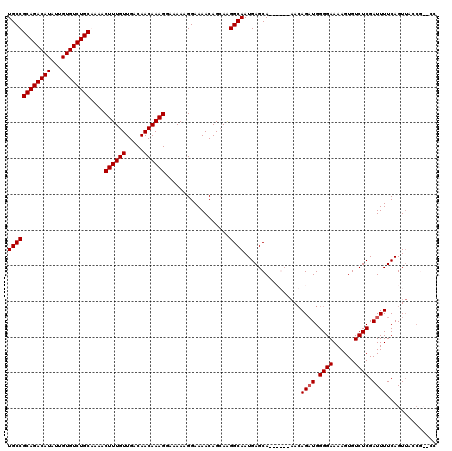
| Location | 7,205,149 – 7,205,269 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.20 |
| Mean single sequence MFE | -28.88 |
| Consensus MFE | -20.67 |
| Energy contribution | -20.68 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.647913 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 7205149 120 - 23771897 GGGGCGGUAACUGAAAAUCGAGACACUUUUCCCCAUCUGUUUGCGUUUGCUCAUUGCCUUCCUGUUUUCCUUUUUCCUUUGUUGUCAACAAAGUUUUGCAGACACAAUAUGUCUGCGGCA ((((.(((((.(((.((.(((((((............))))).)).))..)))))))).)))).....((......((((((.....))))))....(((((((.....))))))))).. ( -29.00) >DroSec_CAF1 18481 118 - 1 GG--AGGUAACUGAAAAUCGAGACACUUUUCCCCAUCUGUUUGCGUUUGCUCAUUGCCUUGCUGUUUUCCUUUUUCCUUUGUUGUCAACAAAGUUUUGCAGACACAAUAUGUCUGCGGCA .(--((((((.(((.((.(((((((............))))).)).))..))))))))))(((.............((((((.....))))))....(((((((.....)))))))))). ( -29.40) >DroEre_CAF1 18933 105 - 1 ---------ACUGAAAAUCGAGACACUUUUCCCCAGCUCUG------UGCUCAUUGCCUUGCUGUUUUCCUUUUCCCUUUGUUGUCAACAAAGUUUCGCAGACACAAUAUGUCUGCGGCA ---------..........(((.(((..............)------)))))..((((..................((((((.....))))))....(((((((.....))))))))))) ( -25.54) >DroYak_CAF1 16087 114 - 1 GGGGCGAGAACUGAAAAUCGAGACACUUUUCCCCAUCUGUU------UGCUCAUUGCCCUGCUGUUUUCCUUUUUGCUUUGUUGUCAACAAAGUUUCGCAGACACAAUAUGUCUGCGGCA (((((((((........))(((..((............)).------..))).)))))))((.............(((((((.....)))))))..((((((((.....)))))))))). ( -31.60) >consensus GG__CGGUAACUGAAAAUCGAGACACUUUUCCCCAUCUGUU______UGCUCAUUGCCUUGCUGUUUUCCUUUUUCCUUUGUUGUCAACAAAGUUUCGCAGACACAAUAUGUCUGCGGCA ...................(((.((......................)))))..((((..................((((((.....))))))....(((((((.....))))))))))) (-20.67 = -20.68 + 0.00)

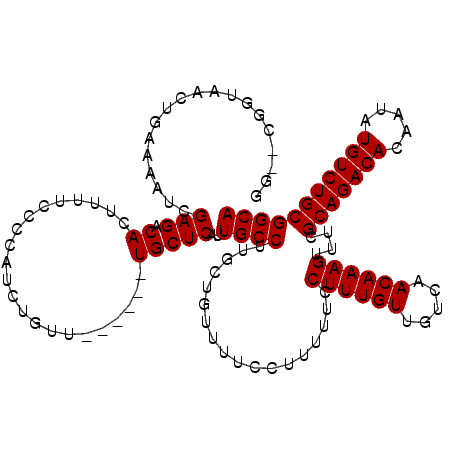

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:36:55 2006