| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 104,696 – 104,789 |
| Length | 93 |
| Max. P | 0.968021 |

| Location | 104,696 – 104,789 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 76.39 |
| Mean single sequence MFE | -17.87 |
| Consensus MFE | -12.25 |
| Energy contribution | -12.68 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.968021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
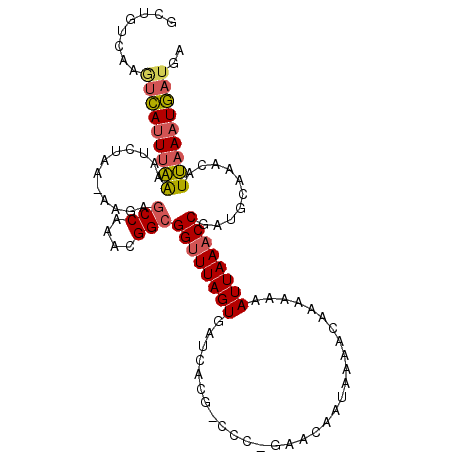
>3L_DroMel_CAF1 104696 93 - 23771897 -------AAGCAUUUAAAAUCUAAAAAGAGCCAAACGGCGGUUUAGUGAUCAU------GAACAAUAAAACAAACAAAUUAAACCGAUGCAAACAUUAAAUGAUGA -------..(((((...............(((....)))((((((((......------..................)))))))))))))...((((....)))). ( -17.16) >DroSec_CAF1 40585 105 - 1 GCUGUCAAGUCAUUUAAAAUCUAAAAAGAGCCAAACGGCGGUUUAGUGAUCACGACCC-GAACAAUAAAACAAAAAAAUUAAACCGAUGCAAACAUUAAAUGAUGA ........(((((((((............(((....)))((((((((...........-..................))))))))..........))))))))).. ( -18.35) >DroSim_CAF1 45843 104 - 1 GCUGUCAAGUCAUUUAAAAUCUAA-AAGAUCCAAACGGCGGUUUAGUGAUCACGACCC-GAACAAUAAAACAAAAAAAUUAAACCGAUGCAAACAUUAAAUGAUGA ........(((((((((.......-.....((....))(((((((((...........-..................))))))))).........))))))))).. ( -15.25) >DroYak_CAF1 57709 104 - 1 ACUUUUAAUUGAGUGGGAGAUUGG-AAAAGCCCAAUGGCGGGUUAGUAAGCAUG-CGUGAAACAAUUAAGCAAAUAAAUUAAGCCAAUACUAACAUCAAAUUAUGA .....(((((..(((..(((((((-...(((((......)))))........((-(.(((.....))).)))...........))))).))..)))..)))))... ( -20.70) >consensus GCUGUCAAGUCAUUUAAAAUCUAA_AAGAGCCAAACGGCGGUUUAGUGAUCACG_CCC_GAACAAUAAAACAAAAAAAUUAAACCGAUGCAAACAUUAAAUGAUGA ........(((((((((............(((....)))((((((((..............................))))))))..........))))))))).. (-12.25 = -12.68 + 0.44)

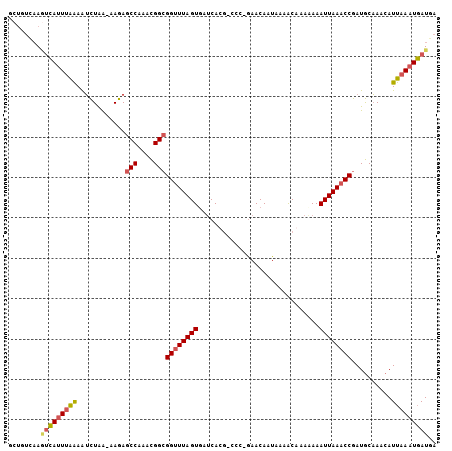
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:30:11 2006