
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 6,845,716 – 6,845,809 |
| Length | 93 |
| Max. P | 0.832413 |

| Location | 6,845,716 – 6,845,809 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 84.43 |
| Mean single sequence MFE | -35.30 |
| Consensus MFE | -25.98 |
| Energy contribution | -26.17 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.832413 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
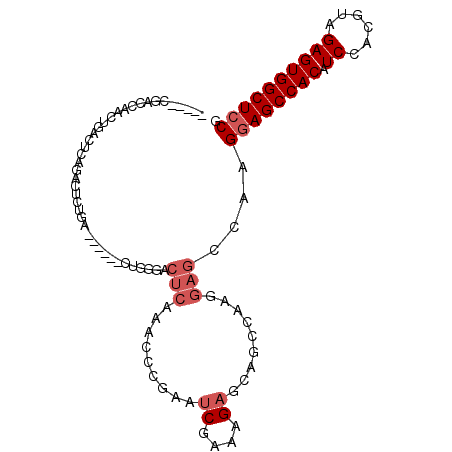
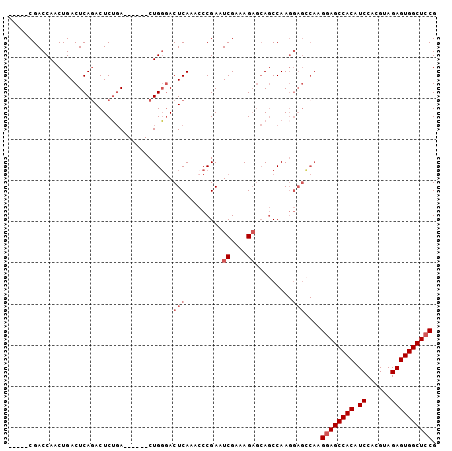
>3L_DroMel_CAF1 6845716 93 + 23771897 CGGAGCCACUCUACGUGGAUGUGGCUUCUUGGCUCCUUGGCUGCUCUUUCGAUUCGGGUUUGAGUGCCAG------UCAGAGUCUGAGUCAGUUGGUCG----- .(((((((((((....))).))))))))..((((..(((((((((((...(((...(((......))).)------)))))))...))))))..)))).----- ( -33.40) >DroSec_CAF1 101912 85 + 1 CGGAGCCACUCUACGUGGAUGUGGCUCC--------UUGGUUGCUCUUUCGAUUCGGUUUUGAGUCCCAG------UCAGAGUCUGAGUCAGUUGGUCG----- .(((((((((((....))).))))))))--------..............(((((((((((((.......------)))))).))))))).........----- ( -30.30) >DroSim_CAF1 106451 93 + 1 CGGAGCCACUCUACGUGGAUGUGGCUCCUUGGCUCCUUGGCUGCUCUUUCGAUUCGGGUUUGAGUCCCAG------UCAGAGUCUGAGUCAGUUGGUCG----- .(((((((((((....))).))))))))..((((..(((((((((((...((((.(((.......)))))------)))))))...))))))..)))).----- ( -38.50) >DroEre_CAF1 118249 101 + 1 CGGAGCCACUCUACGUGGAUGUGGCUCCUUGCCUCCUUGGCU---CUUUCGAUUCGGGUUUGAGUCGCAGUCUGAAUGCGAGUUUGAGUCAGUUGGUCGGAUGG .(((((((((((....))).))))))))..(((...((((((---(.((((((((((..(........)..)))))).))))...)))))))..)))....... ( -39.00) >consensus CGGAGCCACUCUACGUGGAUGUGGCUCCUUGGCUCCUUGGCUGCUCUUUCGAUUCGGGUUUGAGUCCCAG______UCAGAGUCUGAGUCAGUUGGUCG_____ .(((((((((((....))).))))))))..........((((...((...(((((((((((((.............))))).))))))))))..))))...... (-25.98 = -26.17 + 0.19)



| Location | 6,845,716 – 6,845,809 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 84.43 |
| Mean single sequence MFE | -26.65 |
| Consensus MFE | -17.37 |
| Energy contribution | -18.62 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.625216 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 6845716 93 - 23771897 -----CGACCAACUGACUCAGACUCUGA------CUGGCACUCAAACCCGAAUCGAAAGAGCAGCCAAGGAGCCAAGAAGCCACAUCCACGUAGAGUGGCUCCG -----...............(.((((..------.((((..((......)).((....))...)))).)))).)..(.((((((.((......)))))))).). ( -22.90) >DroSec_CAF1 101912 85 - 1 -----CGACCAACUGACUCAGACUCUGA------CUGGGACUCAAAACCGAAUCGAAAGAGCAACCAA--------GGAGCCACAUCCACGUAGAGUGGCUCCG -----...........(((((.......------))))).............((....))........--------((((((((.((......)))))))))). ( -24.20) >DroSim_CAF1 106451 93 - 1 -----CGACCAACUGACUCAGACUCUGA------CUGGGACUCAAACCCGAAUCGAAAGAGCAGCCAAGGAGCCAAGGAGCCACAUCCACGUAGAGUGGCUCCG -----........((.(((.....(((.------.((((.......))))..((....)).))).....))).)).((((((((.((......)))))))))). ( -29.30) >DroEre_CAF1 118249 101 - 1 CCAUCCGACCAACUGACUCAAACUCGCAUUCAGACUGCGACUCAAACCCGAAUCGAAAG---AGCCAAGGAGGCAAGGAGCCACAUCCACGUAGAGUGGCUCCG ((.(((.................(((((.......))))).........(..((....)---)..)..)))))...((((((((.((......)))))))))). ( -30.20) >consensus _____CGACCAACUGACUCAGACUCUGA______CUGGGACUCAAACCCGAAUCGAAAGAGCAGCCAAGGAGCCAAGGAGCCACAUCCACGUAGAGUGGCUCCG ........................................(((.........((....)).........)))....((((((((.((......)))))))))). (-17.37 = -18.62 + 1.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:32:10 2006