| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 6,811,377 – 6,811,471 |
| Length | 94 |
| Max. P | 0.500000 |

| Location | 6,811,377 – 6,811,471 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 85.04 |
| Mean single sequence MFE | -26.58 |
| Consensus MFE | -18.85 |
| Energy contribution | -18.85 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.71 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 6811377 94 - 23771897 AUCUACACCGGAGCAUAUAGUGACACAUUUUCCGCCCCUUGUAGGCGCGAUAUGCGCAUAAUUAAUGAGCCUU-UUAUGGUUUAUUGAACCGCAA--- ..(((((..((.((....((((...))))....))))..)))))((((.....))))....((((((((((..-....)))))))))).......--- ( -26.10) >DroVir_CAF1 100706 83 - 1 --------CGCUGC--AGAGUGACACAUUUUCCGGCCC-UGUAGGCGCGAUAUGCGCAUAAUUAAUGAGCCUU-UUAUGGUUUAUUGAACCGCAA--- --------.((((.--((((((...)))))).))))..-((..(((((.....))))....((((((((((..-....)))))))))).)..)).--- ( -25.40) >DroGri_CAF1 86311 83 - 1 --------CGCAGC--AGAGUGACACAUUUUCCGCCCC-UGUAGGCGCGAUACGCGCAUAAUUAAUGAGCCUU-UUAUGGUUUAUUGAACCGCAA--- --------.((.((--((.(((..........)))..)-)))..(((((...)))))....((((((((((..-....))))))))))...))..--- ( -24.60) >DroWil_CAF1 112145 79 - 1 ------------------AGUGACACAUUUUCCGCCCC-UGUAGGCGCGAUAUGCGCAUAAUUAAUGAGCCUUUUUAUGGUUUAUUGAACCGCACCGC ------------------.(((.(.(((..((((((..-....)))).)).))).))))..((((((((((.......)))))))))).......... ( -20.80) >DroMoj_CAF1 92765 91 - 1 GGCUACAGCGUUGC--AGAGUGACACAUUUUCCGGGCC-UGUAGGCGCGAUAUGCGCAUAAUUAAUGAGCCUU-UUAUGGUUUAUUGAACCGCAA--- ((((((((.((((.--((((((...)))))).).))))-)))))((((.....))))....((((((((((..-....)))))))))).))....--- ( -28.10) >DroAna_CAF1 84143 93 - 1 UUCUACACCGGGGCGCAUAGUGACACAUUUUCCGCCCC-UGUAGGCGCGAUAUGCGCAUAAUUAAUGAGCCUU-UUAUGGUUUAUUGAACCGCAA--- ..(((((..((((((...((((...))))...))))))-)))))((((.....))))....((((((((((..-....)))))))))).......--- ( -34.50) >consensus ________CGCAGC__AGAGUGACACAUUUUCCGCCCC_UGUAGGCGCGAUAUGCGCAUAAUUAAUGAGCCUU_UUAUGGUUUAUUGAACCGCAA___ ...................(((......................((((.....))))....((((((((((.......))))))))))..)))..... (-18.85 = -18.85 + -0.00)

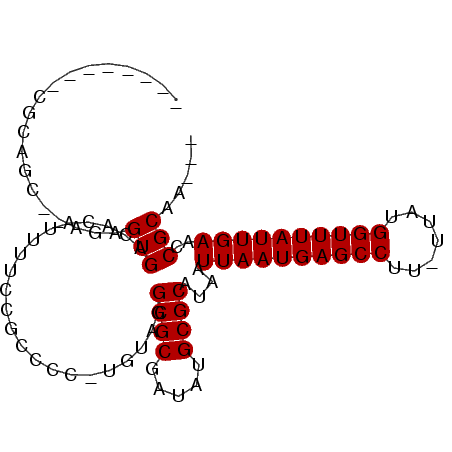
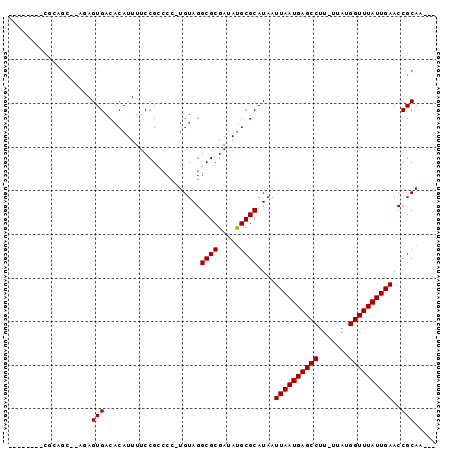
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:32:02 2006