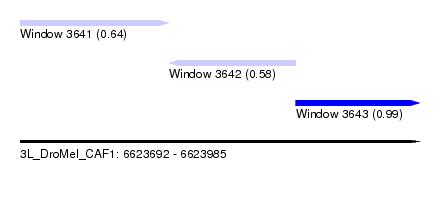
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 6,623,692 – 6,623,985 |
| Length | 293 |
| Max. P | 0.994257 |

| Location | 6,623,692 – 6,623,801 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.48 |
| Mean single sequence MFE | -26.42 |
| Consensus MFE | -21.04 |
| Energy contribution | -21.00 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.639746 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 6623692 109 + 23771897 CCACUAAAU-UCUUU----------GUUAAUGUGCCAUUUAACGGCGCGGACCAGAGAGUCAAAGUCAAGUCGAGUAAUUUCCUUUAAUGGCUUUUCAUUUCAUCGAUAGUGACACUUUC .(((((.((-(((((----------(....((((((.......))))))...)))))))).(((((((((..((....))..))....)))))))............)))))........ ( -25.60) >DroSec_CAF1 731 109 + 1 CCACUAAAU-UCUUU----------GUUAAUGUGCCAUUUAACGGUGCGGACCAGAGAGUCAAAGUCAAGUCGGGUAAUUUCCUUUAAUGGCUUUUCAUUUCAUCGAUAGUGACACUUUC .(((((.((-(((((----------(....((..((.......))..))...)))))))).(((((((....(((......)))....)))))))............)))))........ ( -26.10) >DroSim_CAF1 8866 109 + 1 CCACUAAAU-UCUUU----------GUUAAUGUGCCAUUUAACGGCGCGGACCAGAGAGUCAAAGUCAAGUCGGGUAAUUUCCUUUAAUGGCUUUUCAUUUCAUCGAUAGUGACACUUUC .(((((.((-(((((----------(....((((((.......))))))...)))))))).(((((((....(((......)))....)))))))............)))))........ ( -28.50) >DroYak_CAF1 2704 110 + 1 CCACUAAAUUUCUUU----------GCUAAUGUGCCAUUUAACGGCGCGGACCAGAGACUCAAAGUCAAGUCGGGUAAUUUCCUUUAAUGGCUUUUCAUUUCAUCGAUAGUGACACUUUC .(((((....(((((----------(....((((((.......))))))...))))))...(((((((....(((......)))....)))))))............)))))........ ( -26.80) >DroAna_CAF1 839 119 + 1 CUACUAAAU-UUUUUCCUCUAUGUCGUUAAUGUGCCAUUUAACGGCGGAGCGAGCAAAGUCAAAGUCAAGUUGGGUAAUUUCCUCUAAUGGCUUUUCAUUUCAUCGAUAGUGACACUUUC .(((((...-......((((..((((((((........))))))))))))(((..(((...(((((((....(((......)))....)))))))...)))..))).)))))........ ( -25.10) >consensus CCACUAAAU_UCUUU__________GUUAAUGUGCCAUUUAACGGCGCGGACCAGAGAGUCAAAGUCAAGUCGGGUAAUUUCCUUUAAUGGCUUUUCAUUUCAUCGAUAGUGACACUUUC .(((((........................((((((.......)))))).....((((...(((((((....(((......)))....)))))))...)))).....)))))........ (-21.04 = -21.00 + -0.04)

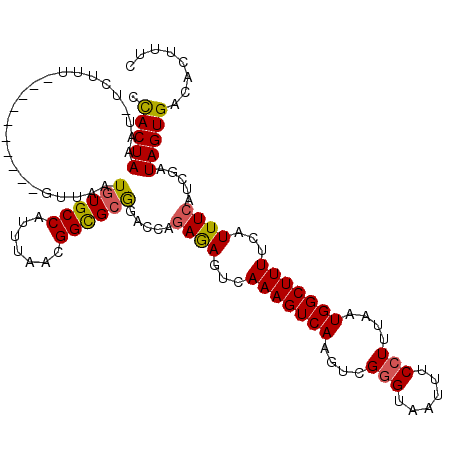
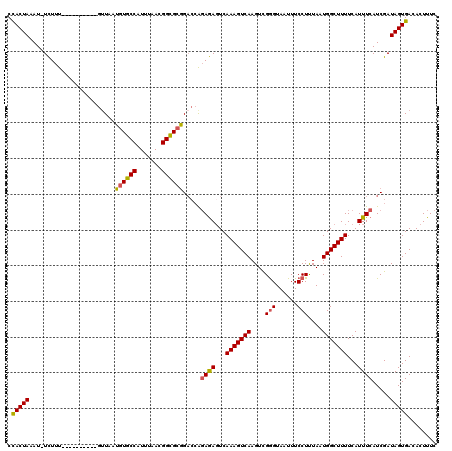
| Location | 6,623,801 – 6,623,894 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 95.04 |
| Mean single sequence MFE | -30.18 |
| Consensus MFE | -24.79 |
| Energy contribution | -24.48 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.575528 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 6623801 93 - 23771897 UUAGCACUUCUUUUUCGGAAGGGUGCGGGG--GCUUAGUAAUUGACCACGCCCCCGUUGUGUAGUUGUCUUUUUUACGGCAUCACGAUGGCCAUU ...(((((.((((....)))))))))((((--((...((.....))...))))))((((((....((((........)))).))))))....... ( -30.70) >DroSec_CAF1 840 93 - 1 UUAGCACUUCUUUUUCGGAAGGGUGCUGGG--GCUUAGUAAUUGACCACGCCCCCGUUGUGUAGUUGUCUUUUUUACGGCAUCACGAUGGCCAUU .(((((((.((((....)))))))))))((--((...((.....))...))))((((((((....((((........)))).))))))))..... ( -29.50) >DroSim_CAF1 8975 93 - 1 UUAGCACUUAUUUUUCGGAAGGGUGCGGGG--GCUUAGUAAUUGACCACGCCCCCGUUGUGUAGUUGUCUUUUUUACGGCAUCACGAUGGCCAUU ...(((((..((.....))..)))))((((--((...((.....))...))))))((((((....((((........)))).))))))....... ( -29.00) >DroYak_CAF1 2814 95 - 1 UUAGCACUUAUUUUUCGGAAGGGUGCGGGGCGGUGUAGUAAUUGACCACGCCCCCGUCUUGUAGUUGUCUUUUUUACGGCAUCAGGAUGGCCAUU ...(((((..((.....))..)))))((((((..((........))..))))))(((((((....((((........)))).)))))))...... ( -31.50) >consensus UUAGCACUUAUUUUUCGGAAGGGUGCGGGG__GCUUAGUAAUUGACCACGCCCCCGUUGUGUAGUUGUCUUUUUUACGGCAUCACGAUGGCCAUU ................((..(((((.((.................)).)))))((((((((....((((........)))).))))))))))... (-24.79 = -24.48 + -0.31)


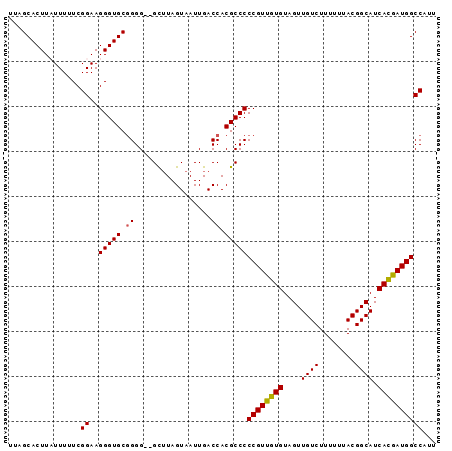
| Location | 6,623,894 – 6,623,985 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.19 |
| Mean single sequence MFE | -24.10 |
| Consensus MFE | -22.88 |
| Energy contribution | -22.56 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.95 |
| SVM decision value | 2.46 |
| SVM RNA-class probability | 0.994257 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 6623894 91 + 23771897 ACAGGAAAUGAUUUCCUGGCCACU-----------------------------CUCAACCCUCUGGUAAUGAAAUCAUGAAUUAGAUUUUGAGCUUGAAAAUUGUUAUUGUUUUCGGCCA ..(((((.....)))))((((...-----------------------------(((((...((((((.(((....)))..))))))..)))))...((((((.......)))))))))). ( -25.00) >DroSec_CAF1 933 91 + 1 ACAGGAAAUGAUUUUCUGGCCACU-----------------------------CUCAACCCUCUGGUAAUGAAAUCAUGAAUUAGAUUUUGAGCUUGAAAAUUGUUAUUGUUUUCGGCCA ...(((((....)))))((((...-----------------------------(((((...((((((.(((....)))..))))))..)))))...((((((.......)))))))))). ( -23.40) >DroSim_CAF1 9068 91 + 1 ACAGGAAAUGAUUUUCUGGCCACU-----------------------------CUCAACCCUCUGGUAAUGAAAUCAUGAAUUAGAUUUUGAGCUUGAAAAUUGUUAUUGUUUUCGGCCA ...(((((....)))))((((...-----------------------------(((((...((((((.(((....)))..))))))..)))))...((((((.......)))))))))). ( -23.40) >DroYak_CAF1 2909 95 + 1 ACAGGAAAUGAUUUUCUGGCCACUCU-------------------------CUCUCAACCCUCUGGUAAUGAAAUCAUGAAUUAGAUUUUGAGCUUGAAAAUUGUUAUUGUUUUCGGCCA ...(((((....)))))((((.....-------------------------..(((((...((((((.(((....)))..))))))..)))))...((((((.......)))))))))). ( -23.40) >DroAna_CAF1 1058 120 + 1 ACAGGAAAUGAUUUUCUGGCCACUCCUUAGCCUCCAGCCCUCAGCCCUUAGCUCUCAGCCCUCUGGUAAUGAAAUCAUGAAUUAGAUUUUGAGCUUGAAAAUUGUUAUUGUUUUCGGCCA ...(((((....)))))((((............................(((((...(((....)))...((((((........))))))))))).((((((.......)))))))))). ( -25.30) >consensus ACAGGAAAUGAUUUUCUGGCCACU_____________________________CUCAACCCUCUGGUAAUGAAAUCAUGAAUUAGAUUUUGAGCUUGAAAAUUGUUAUUGUUUUCGGCCA ..(((((.....)))))((((................................(((((...((((((.(((....)))..))))))..)))))...((((((.......)))))))))). (-22.88 = -22.56 + -0.32)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:30:26 2006