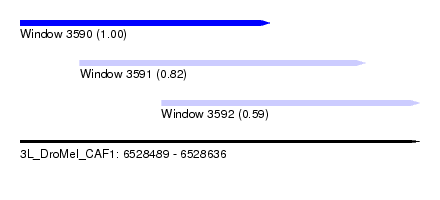

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 6,528,489 – 6,528,636 |
| Length | 147 |
| Max. P | 0.999626 |

| Location | 6,528,489 – 6,528,581 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 96.41 |
| Mean single sequence MFE | -23.88 |
| Consensus MFE | -22.08 |
| Energy contribution | -22.32 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.92 |
| SVM decision value | 3.80 |
| SVM RNA-class probability | 0.999626 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
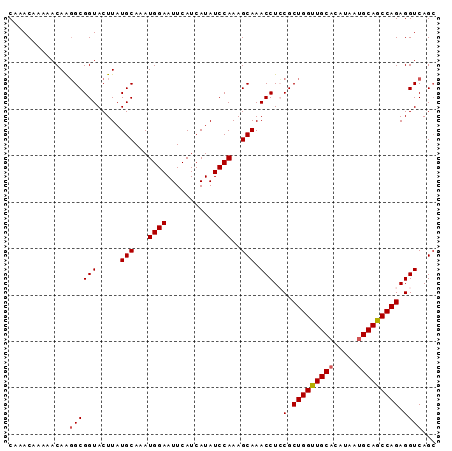
>3L_DroMel_CAF1 6528489 92 + 23771897 CAAACAAAAACAAGGCGGUACUUAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGC .............((.(((.....(((...((((..........))))..))).))).)).(((((((((.....)))))))))........ ( -23.40) >DroSec_CAF1 19755 92 + 1 CAAACAAAAACAAGGCGGUACUUAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGC .............((.(((.....(((...((((..........))))..))).))).)).(((((((((.....)))))))))........ ( -23.40) >DroSim_CAF1 19971 92 + 1 CAAACAAAAACAAGGCGGUACUUAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGG ...........(((......))).(((...((((..........))))..)))..(((((.(((((((((.....))))))))).))..))) ( -23.90) >DroEre_CAF1 19904 92 + 1 CAAACAAAAACAAGGCGGUACUUAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUGCGCUGGCUGCACUUAAUGCAGCCAGAGGUCAGC .............((((((.....(((...((((..........))))..))).)))..(.(((((((((.....))))))))).))))... ( -25.40) >DroYak_CAF1 20223 92 + 1 CAAACAAAAACAGCGCGGUACUUAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUGCGCUGGUUGCCCGUAAUGCAGCCAGAGGUCAGC ...........(((((((......(((...((((..........))))..)))...)))))))((((((.......)))))).......... ( -23.30) >consensus CAAACAAAAACAAGGCGGUACUUAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGC .............((((((.....(((...((((..........))))..))).)))..(.(((((((((.....))))))))).))))... (-22.08 = -22.32 + 0.24)


| Location | 6,528,511 – 6,528,616 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 92.29 |
| Mean single sequence MFE | -32.70 |
| Consensus MFE | -25.84 |
| Energy contribution | -27.12 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824384 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
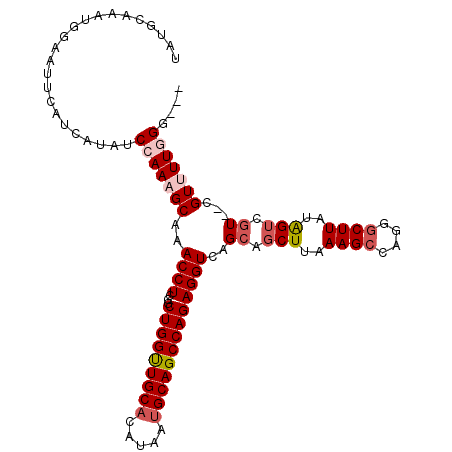
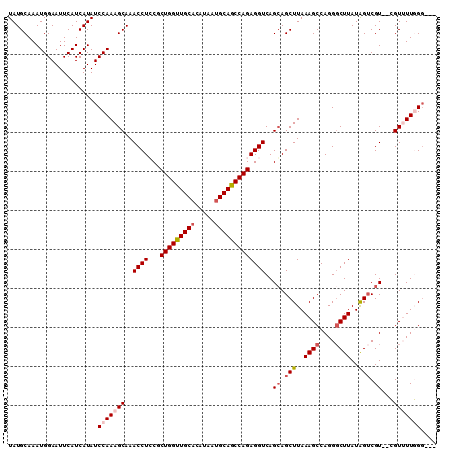
>3L_DroMel_CAF1 6528511 105 + 23771897 UAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGUCUUAUGGUCGU--CGUUUUGGG--- ...((...((((..........))))..((..(((((...((((((((.....)))))))))))))..)).))(((((((..(..(....)..)..--.))))))).--- ( -30.80) >DroSec_CAF1 19777 105 + 1 UAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGGCUUAUAGUCGU--CGUUUUGGG--- .......................(((((((......((.(((((((((.....))))))))).))...((.(((..((((....))))..))).))--.))))))).--- ( -32.60) >DroSim_CAF1 19993 105 + 1 UAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGGAGCUUAAAGCCAGGGCUUAUAGUCGU--CGUUUUGGG--- .......................(((((((...(((((.(((((((((.....))))))))).))..))).(((..((((....))))..)))...--.))))))).--- ( -34.20) >DroEre_CAF1 19926 107 + 1 UAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUGCGCUGGCUGCACUUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGGCUUAUAGACGUGUGGUGUUUGU--- ...(((((.......((((((((.....((..((((...(((((((((.....)))))))))))))..)).(((..((((....))))..)).))))))))))))))--- ( -34.31) >DroYak_CAF1 20245 110 + 1 UAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUGCGCUGGUUGCCCGUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGGCUUAUAGUCGUGUGGUGUUUGGUUC ......(((.(((..((((((((.....((..((((...((((((((.......))))))))))))..)).(((..((((....))))..))).))))))))))).))). ( -31.60) >consensus UAUGCAAAUGGAAUUCAUCAUAUCCAAAGCAAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGGCUUAUAGUCGU__CGUUUUGGG___ .......................(((((((..((((...(((((((((.....)))))))))))))..((.(((..((((....))))..))).))...))))))).... (-25.84 = -27.12 + 1.28)

| Location | 6,528,541 – 6,528,636 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 86.99 |
| Mean single sequence MFE | -31.28 |
| Consensus MFE | -21.26 |
| Energy contribution | -22.14 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.594110 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
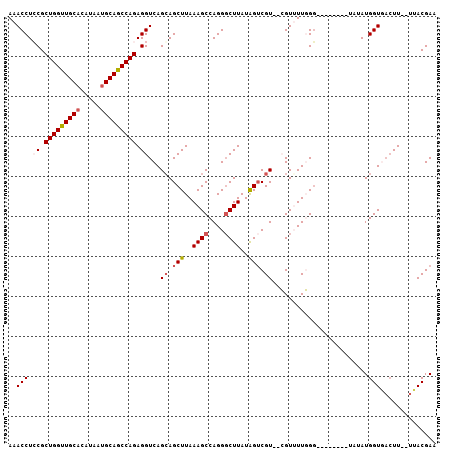
>3L_DroMel_CAF1 6528541 95 + 23771897 AAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGUCUUAUGGUCGU--CGUUUUGGG--------UAUAUGGUGACUU--UUACGAA .......(((((((((((.....)))))))))((((((.(((((((((((..(..(....)..)..--.))))))))--------)...)).))))))--...)).. ( -32.70) >DroSec_CAF1 19807 95 + 1 AAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGGCUUAUAGUCGU--CGUUUUGGG--------UAUAUGGUGACUU--UUACGAA .......(((((((((((.....)))))))))((((((.(((((((((((.(.(((.....))).)--.))))))))--------)...)).))))))--...)).. ( -33.40) >DroSim_CAF1 20023 95 + 1 AAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGGAGCUUAAAGCCAGGGCUUAUAGUCGU--CGUUUUGGG--------UAUAUGGUGACUU--UUACGAA ...(((((.(((((((((.....))))))))).))..))).(((((((((.(.(((.....))).)--.))))))))--------)....(....)..--....... ( -32.30) >DroEre_CAF1 19956 98 + 1 AAACCUGCGCUGGCUGCACUUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGGCUUAUAGACGUGUGGUGUUUGU--------UAUAUGGUAACUU-UUUACGAA ....((((.(((((((((.....)))))))))......))))....((((....)))).....(((((((......)--------))))))((((...-.))))... ( -29.30) >DroYak_CAF1 20275 107 + 1 AAACCUGCGCUGGUUGCCCGUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGGCUUAUAGUCGUGUGGUGUUUGGUUCAUACAUAUAUGGUACAUUUUUAACGAA ....((((.((((((((.......))))))))......))))......((((.(((.....)))((((((........)))))).....)))).............. ( -28.70) >consensus AAACCUCCGCUGGUUGCACAUAAUGCAGCCAGAGGUCAGCAGCUUAAAGCCAGGGCUUAUAGUCGU__CGUUUUGGG________UAUAUGGUGACUU__UUACGAA ..(((.((.(((((((((.....))))))))).))...((.(((..((((....))))..))).))........................))).............. (-21.26 = -22.14 + 0.88)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:29:37 2006