
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 6,478,364 – 6,478,459 |
| Length | 95 |
| Max. P | 0.968336 |

| Location | 6,478,364 – 6,478,459 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 80.09 |
| Mean single sequence MFE | -26.15 |
| Consensus MFE | -17.32 |
| Energy contribution | -19.08 |
| Covariance contribution | 1.77 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.94 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961970 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
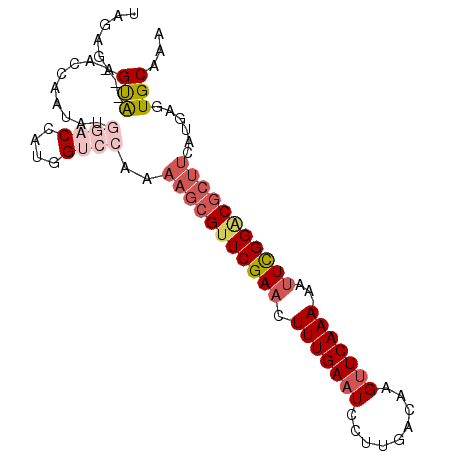
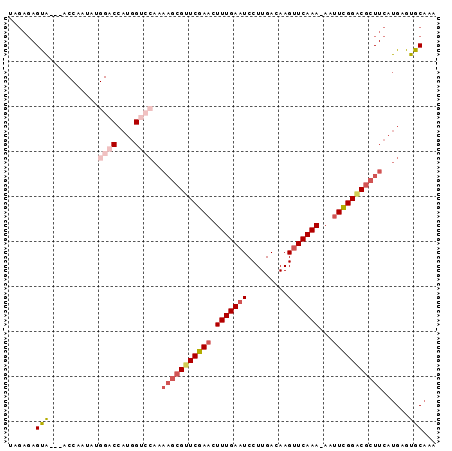
>3L_DroMel_CAF1 6478364 95 + 23771897 UAGAGAGUAUGGACCAGUGCGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGCUCAAAAAAAUUGGACGCUUCAUGAGUGCAAA ......((((((((((..........))))))..((((((((((..(((((.............)))))....)))))))))).....))))... ( -24.32) >DroSec_CAF1 12909 84 + 1 UAGAGAGUAUGGA----------CCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAA-AAUUCGGACGCUUCAUGAGUGCAAA ......(((((((----------(....))))..(((((((((((.(((((((.........)))))))-..))))))))))).....))))... ( -27.60) >DroSim_CAF1 12999 83 + 1 UAGAGAGUAUGGA----------CCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAA-AAUUCGGGCGCUUCAUGAGUGCAA- ......(((((((----------(....))))..(((((((((((.(((((((.........)))))))-..))))))))))).....))))..- ( -27.10) >DroEre_CAF1 13520 91 + 1 UAGAGAGCG---ACCAAUAUGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAA-AAUUCGGACGCUUCAUGAGUGCAAA ......(((---..((...(((((....))))).(((((((((((.(((((((.........)))))))-..)))))))))))..))..)))... ( -30.10) >DroYak_CAF1 13485 91 + 1 UAGAGAGCA---ACCUAUAUGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAA-UAUUCGGACGCUUCAUGAGUGCAAA ......(((---..(....(((((....))))).(((((((((((.(((((((.........)))))))-..)))))))))))...)..)))... ( -29.10) >DroAna_CAF1 14147 74 + 1 UGGUG-------ACCAAUAUGGACCAUGGUCCG-AAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAA-AAUUCGGCCA------------GAA ((((.-------.((.....))))))(((.(((-((...((.((((((............)))))).))-..))))))))------------... ( -18.70) >consensus UAGAGAGUA___ACCAAUAUGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAA_AAUUCGGACGCUUCAUGAGUGCAAA ......(((...........((((....))))..(((((((((((.(((((((.........)))))))...)))))))))))......)))... (-17.32 = -19.08 + 1.77)

| Location | 6,478,364 – 6,478,459 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 80.09 |
| Mean single sequence MFE | -26.13 |
| Consensus MFE | -16.64 |
| Energy contribution | -18.57 |
| Covariance contribution | 1.93 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.13 |
| Structure conservation index | 0.64 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.968336 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
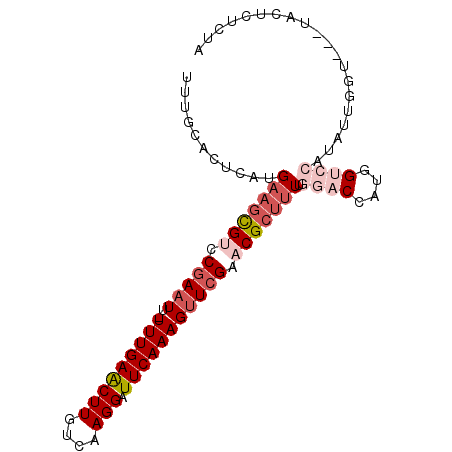
>3L_DroMel_CAF1 6478364 95 - 23771897 UUUGCACUCAUGAAGCGUCCAAUUUUUUUGAGCUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCGCACUGGUCCAUACUCUCUA ...........(((((((.(.....((((((((((....))).)))))))...).)))))))(((((((((.....)).)))))))......... ( -23.30) >DroSec_CAF1 12909 84 - 1 UUUGCACUCAUGAAGCGUCCGAAUU-UUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGG----------UCCAUACUCUCUA ...........(((((((.((((((-((.((((((....))).))))))))))).)))))))(((((....)----------))))......... ( -26.60) >DroSim_CAF1 12999 83 - 1 -UUGCACUCAUGAAGCGCCCGAAUU-UUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGG----------UCCAUACUCUCUA -..........((((((..((((((-((.((((((....))).)))))))))))..))))))(((((....)----------))))......... ( -24.60) >DroEre_CAF1 13520 91 - 1 UUUGCACUCAUGAAGCGUCCGAAUU-UUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCAUAUUGGU---CGCUCUCUA ...(((((.(((((((((.((((((-((.((((((....))).))))))))))).)))))).(((((....)))))))).)))---.))...... ( -29.10) >DroYak_CAF1 13485 91 - 1 UUUGCACUCAUGAAGCGUCCGAAUA-UUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCAUAUAGGU---UGCUCUCUA ...(((..(..(((((((.(((((.-(((((((((....))).))))))))))).)))))))(((((....)))))....)..---)))...... ( -30.50) >DroAna_CAF1 14147 74 - 1 UUC------------UGGCCGAAUU-UUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUU-CGGACCAUGGUCCAUAUUGGU-------CACCA ...------------((((((((((-(((((.....))))))))))...............-.((((....)))).....)))-------))... ( -22.70) >consensus UUUGCACUCAUGAAGCGUCCGAAUU_UUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCAUAUUGGU___UACUCUCUA ...........(((((((.(((((..(((((((((....))).))))))))))).))))))).((((....)))).................... (-16.64 = -18.57 + 1.93)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:29:05 2006