
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,832,194 – 5,832,346 |
| Length | 152 |
| Max. P | 0.939507 |

| Location | 5,832,194 – 5,832,311 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 79.57 |
| Mean single sequence MFE | -24.60 |
| Consensus MFE | -19.67 |
| Energy contribution | -20.50 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.30 |
| SVM RNA-class probability | 0.939507 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5832194 117 + 23771897 GCAACUCGGAUCGGCGGGACAGCCGGCAGAAAUCACGUUAAGCGCAAUAAAAAGCGCGAAACAUUUUAACGCAUCCCGGCGAUAAAGUAUCUGAAAACAUUGCCCCAGACAAUGUUC- .....((((((((((......))))...........(((..((((........))))..)))((((((.(((......))).)))))))))))).(((((((.......))))))).- ( -31.80) >DroVir_CAF1 46715 100 + 1 AGC---------A--GCA-----AAACAGAAAUCACGUUAAGCGCAAUAAAAAGCGCGAAACAUUUUAACGCGUCGCGGCGAUAAAGUAUCUGAAAACAUUGCC-CCGACAAUGUCC- ...---------.--...-----...((((......(((..((((........))))..)))((((((.(((......))).)))))).))))...((((((..-....))))))..- ( -23.90) >DroGri_CAF1 50897 110 + 1 AGC----GAGCCG--AGAACAGAAAACAGAAAUCACGUUAAGCGCAAUAAAAAGCGCGAAACAUUUUAACGCGUCGAGGCGAUAAAGUAUCUGAAAACAUUGCC-CCGACAAUGUCC- .((----(....(--(...(........)...))..(((..((((........))))..))).......)))((((.((((((...((........))))))))-.)))).......- ( -25.20) >DroYak_CAF1 43594 92 + 1 -------------------------GCAGAAAUCACGUUAAGCGCAAUAAAAAGCGCGAAACAUUUUAACGCAUCCCGGCGAUAAAGUAUCUGAAAACAUUGCCCCAGACAAUGUUC- -------------------------.((((......(((..((((........))))..)))((((((.(((......))).)))))).))))..(((((((.......))))))).- ( -23.80) >DroAna_CAF1 43060 105 + 1 --AA--CAGAUCCA---UACAGCUGGCUGAAAUCACGUUAAGCUCAAUAAAAAGCGCGAAACAUUUUAACGCAUCCCGGCGAUAAAGUAUCUGAAAACAUUGCCCCAGACAA------ --..--((((((..---((..(((((...............(((........)))(((...........)))...)))))..))..).)))))...................------ ( -17.10) >DroPer_CAF1 44340 104 + 1 ACA----GGGUC-------CAG--AACAGAAAUCACGUUAAGCGCAAUAAAAAGCGCGAAACAUUUUAACGCAUCUCAGCGAUAAAGUAUCUGAAAACAUUGCC-CAGACAAUGUCCA ...----((...-------...--..((((......(((..((((........))))..)))((((((.(((......))).)))))).))))...((((((..-....)))))))). ( -25.80) >consensus A_A____GG_UC_______CAG__AACAGAAAUCACGUUAAGCGCAAUAAAAAGCGCGAAACAUUUUAACGCAUCCCGGCGAUAAAGUAUCUGAAAACAUUGCC_CAGACAAUGUCC_ ..........................((((......(((..((((........))))..)))((((((.(((......))).)))))).))))...((((((.......))))))... (-19.67 = -20.50 + 0.83)


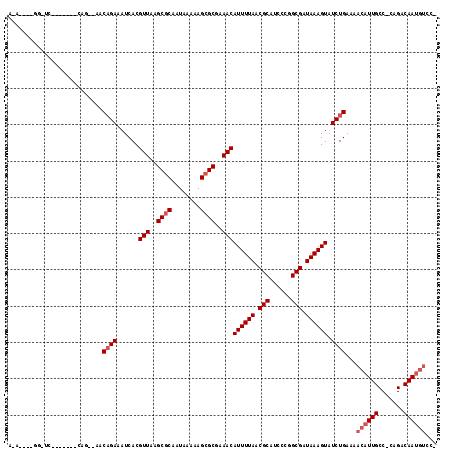
| Location | 5,832,194 – 5,832,311 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 79.57 |
| Mean single sequence MFE | -28.93 |
| Consensus MFE | -22.32 |
| Energy contribution | -23.32 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.833723 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5832194 117 - 23771897 -GAACAUUGUCUGGGGCAAUGUUUUCAGAUACUUUAUCGCCGGGAUGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCUUAACGUGAUUUCUGCCGGCUGUCCCGCCGAUCCGAGUUGC -(((((((((.....)))))))))......((((.((((.(((((((((..((.(((((..((((........))))..)))))..))..))).....)))))).))))..))))... ( -36.20) >DroVir_CAF1 46715 100 - 1 -GGACAUUGUCGG-GGCAAUGUUUUCAGAUACUUUAUCGCCGCGACGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCUUAACGUGAUUUCUGUUU-----UGC--U---------GCU -(((((((((...-.)))))))))...((((...))))((.((((.(((..((.(((((..((((........))))..)))))..))..))).)-----)))--.---------)). ( -28.80) >DroGri_CAF1 50897 110 - 1 -GGACAUUGUCGG-GGCAAUGUUUUCAGAUACUUUAUCGCCUCGACGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCUUAACGUGAUUUCUGUUUUCUGUUCU--CGGCUC----GCU -(((((..(((((-(((..((....))((((...))))))))))))(((((((........((((........))))))))))).............))))).--......----... ( -30.50) >DroYak_CAF1 43594 92 - 1 -GAACAUUGUCUGGGGCAAUGUUUUCAGAUACUUUAUCGCCGGGAUGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCUUAACGUGAUUUCUGC------------------------- -(((((((((.....))))))))).((((...((((.(((......))).))))(((((..((((........))))..)))))....)))).------------------------- ( -26.10) >DroAna_CAF1 43060 105 - 1 ------UUGUCUGGGGCAAUGUUUUCAGAUACUUUAUCGCCGGGAUGCGUUAAAAUGUUUCGCGCUUUUUAUUGAGCUUAACGUGAUUUCAGCCAGCUGUA---UGGAUCUG--UU-- ------.((((((..(......)..))))))....(((((.(.((.((((....)))).)).)((((......)))).....)))))..((((((......---)))..)))--..-- ( -22.50) >DroPer_CAF1 44340 104 - 1 UGGACAUUGUCUG-GGCAAUGUUUUCAGAUACUUUAUCGCUGAGAUGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCUUAACGUGAUUUCUGUU--CUG-------GACCC----UGU ..((((((((...-.))))))))((((((.((((((.(((......))).))))(((((..((((........))))..))))).......)))--)))-------))...----... ( -29.50) >consensus _GGACAUUGUCUG_GGCAAUGUUUUCAGAUACUUUAUCGCCGGGAUGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCUUAACGUGAUUUCUGCU__CUG_______GA_CC____U_U .(((((((((.....))))))))).((((...((((.(((......))).))))(((((..((((........))))..)))))....)))).......................... (-22.32 = -23.32 + 1.00)


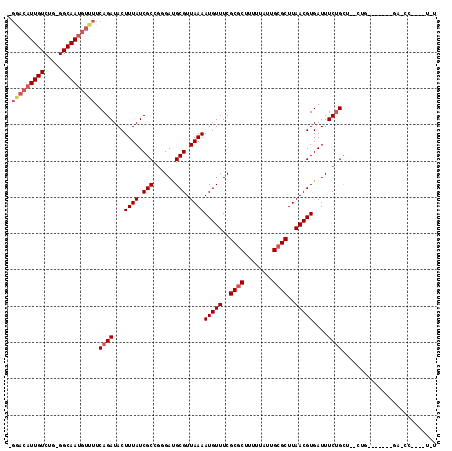
| Location | 5,832,234 – 5,832,346 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.54 |
| Mean single sequence MFE | -36.65 |
| Consensus MFE | -24.04 |
| Energy contribution | -25.07 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5832234 112 - 23771897 AUUCGCGUCGAG--GCUUAGCGCCUUGAAAUCGCCGG----GAACAUUGUCUG--GGGCAAUGUUUUCAGAUACUUUAUCGCCGGGAUGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCU ...(((.(((((--((.....))))))).(((.((((----(((((((((...--..)))))))))...((((...)))).))))))))))............((((........)))). ( -40.00) >DroVir_CAF1 46739 106 - 1 AUUCGCGUCGAGGGGCUUAGCGCCCC--AAUU---------GGACAUUGUCGG---GGCAAUGUUUUCAGAUACUUUAUCGCCGCGACGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCU ...(((((((.(((((.....)))))--....---------(((((((((...---.))))))).....((((...)))).)).)))))))............((((........)))). ( -39.90) >DroPse_CAF1 44319 115 - 1 AUUCGCGUUGAG--GCUUAGCGCCUGGGAAUGGGCUGUUCUGGACAUUGUCUG---GGCAAUGUUUUCAGAUACUUUAUCGCUGAGAUGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCU ...((((((..(--((.....((((......)))).((((((((((((((...---.))))))))..)))).))......)))..))))))............((((........)))). ( -37.10) >DroGri_CAF1 50931 106 - 1 AUUCGCGUCGAGGGGCUUAGCGCCUC--AAUU---------GGACAUUGUCGG---GGCAAUGUUUUCAGAUACUUUAUCGCCUCGACGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCU ...(((((((((((((.....)))((--....---------(((((((((...---.)))))))))...))..........))))))))))............((((........)))). ( -40.10) >DroMoj_CAF1 48780 109 - 1 AUUCCCGUCCAGGGGCUUAGCGCCCC--AAUU---------GGACAUUGUCGGUUCGAAAAUGUUUUCAGAUACUUUAUCGCAGCGACGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCU ......((((((((((.....)))))--...)---------))))...((((.(.(((.((.((........)).)).))).).))))...............((((........)))). ( -33.40) >DroAna_CAF1 43093 100 - 1 AUUCGCGUUGAG--GCUUAGCGCCUUGGAA----------------UUGUCUG--GGGCAAUGUUUUCAGAUACUUUAUCGCCGGGAUGCGUUAAAAUGUUUCGCGCUUUUUAUUGAGCU ...((((((..(--((.....((((..((.----------------...))..--))))..........((((...)))))))..))))))..............((((......)))). ( -29.40) >consensus AUUCGCGUCGAG__GCUUAGCGCCUC__AAUU_________GGACAUUGUCGG___GGCAAUGUUUUCAGAUACUUUAUCGCCGCGACGCGUUAAAAUGUUUCGCGCUUUUUAUUGCGCU ....(((.(((.......(((((..................((((((((((.....))))))))))..(((((.((((.(((......))).)))).))))).))))).....)))))). (-24.04 = -25.07 + 1.03)


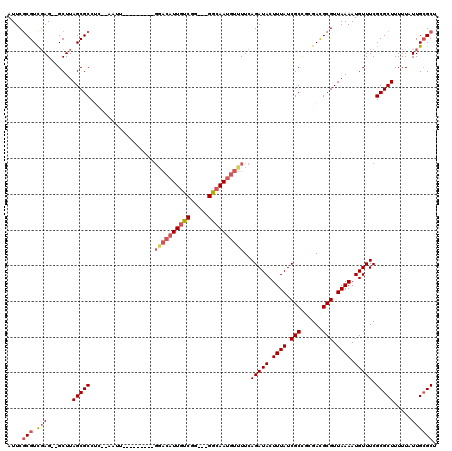
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:23 2006