
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 597,060 – 597,155 |
| Length | 95 |
| Max. P | 0.713999 |

| Location | 597,060 – 597,155 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.75 |
| Mean single sequence MFE | -26.48 |
| Consensus MFE | -13.54 |
| Energy contribution | -13.79 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.713999 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
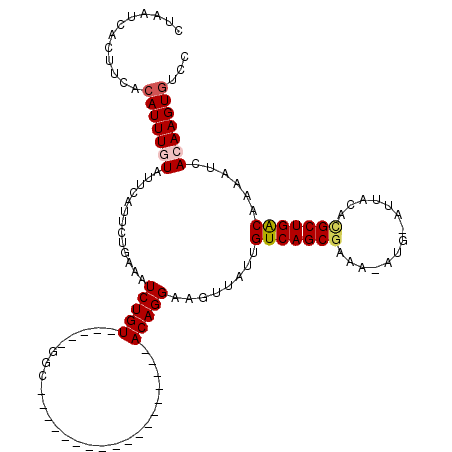
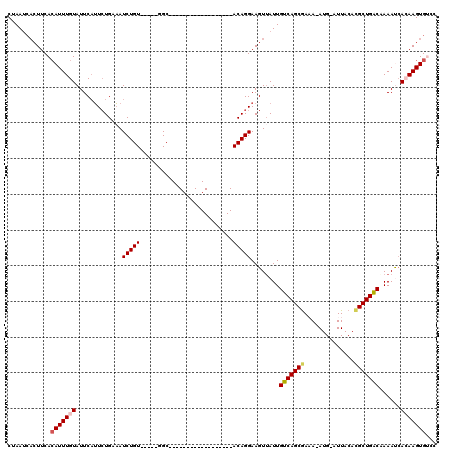
>3L_DroMel_CAF1 597060 95 - 23771897 CCAAUCACUUCAAAUUUGUAUUCAUUCUGAAAUCUGU-----GGC------------------ACAGGAAGUGAUUGUCAGCGAAA-AUG-AUUACACGCUGGCCAAAUGACAAGUGCCC ..(((((((((.........(((.....)))..(((.-----...------------------.))))))))))))(((((((...-...-......)))))))................ ( -22.90) >DroPse_CAF1 15432 113 - 1 CUAAUCACUUCACAUUUGUAUUCAUUCUGUAAUCUGU-----GGCACAGCACCAGCCACAAGCACAGGAAGUUAUUGUCAGCGAAA-AUG-AUUACAUGCUGACAAAACCAGAAGUGUCC .....((((((.............((((((....(((-----(((.........))))))...)))))).(((.((((((((....-(((-....))))))))))))))..))))))... ( -32.60) >DroEre_CAF1 10228 94 - 1 CUAAUCACUUCACAUUUGUAUUCAUUCUGAAAUCUGU-----GGC------------------ACAGGAAGUUAUUGUCAGCGA-A-AUG-AUUACACGCUGGCCAAAUCACAAGUGGCC ............(((((((.(((.....))).....(-----(((------------------((((.......))))(((((.-.-...-......)))))))))....)))))))... ( -19.20) >DroYak_CAF1 10564 95 - 1 CUAA----UUCACAUUUGUAUUCAUUCUGAAAUCUGUGGUGUGGC------------------ACAGGAAGUUAUUGUCAGCGA-A-AUG-AUUACACGCUGGCCAAAUCACAAGUGUCC ....----...((((((((.............((((((......)------------------)))))........(((((((.-.-...-......)))))))......)))))))).. ( -23.20) >DroAna_CAF1 14723 95 - 1 CUAAUCACUUCACAUUUGUAUUCCGUGUGUUAUCUGU-----GGC------------------ACAGGAAGUUAUUGUCAGCAAAA-AUG-AUUACACGCUGACAAAAUCACAAGUGUCC ...........((((((((.((((.(((((((....)-----)))------------------)))))))....((((((((....-...-.......))))))))....)))))))).. ( -28.64) >DroPer_CAF1 16019 115 - 1 CUAAUCACUUCACAUUUGUAUUCAUUCUGUAAUCUGU-----GGCACAGCACCAGCCACAAGCACAGGAAGUUAUUGUCAGCGAAAAAUGAAUUACAUGCUGACAAAACCAGAAGUGUCC .....((((((.............((((((....(((-----(((.........))))))...)))))).(((.(((((((((..............))))))))))))..))))))... ( -32.34) >consensus CUAAUCACUUCACAUUUGUAUUCAUUCUGAAAUCUGU_____GGC__________________ACAGGAAGUUAUUGUCAGCGAAA_AUG_AUUACACGCUGACAAAAUCACAAGUGUCC ............(((((((.............(((((..........................)))))........(((((((..............)))))))......)))))))... (-13.54 = -13.79 + 0.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:36:28 2006