
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,583,677 – 5,583,787 |
| Length | 110 |
| Max. P | 0.888410 |

| Location | 5,583,677 – 5,583,787 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 86.50 |
| Mean single sequence MFE | -28.80 |
| Consensus MFE | -18.26 |
| Energy contribution | -19.32 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.644374 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5583677 110 + 23771897 U-UAG--UUUUUAGUUUGUU-GCUUACUGCAACUGUUUUUGGUCAGUGCUGCCAUUGGAUGGCAGCUUACUAUAAUUUUUAUGAUGAAUUGCCUGGCCGCGUGUGAAAUCUGCA .-(((--.(((((....(((-((.....))))).......((((((.((((((((...))))))))......((((((.......)))))).)))))).....))))).))).. ( -30.90) >DroSim_CAF1 2236 105 + 1 --------UUAUAUAUUUUU-GUUUACUGCAACUGUUUUUGGUCAGUGCUGCCAUUGGAUGGCAGCUUACUAUAAUUUUUAUGAUGAAUUGCUUGGCCGCGUGUGAAAUCUGCA --------((((((....((-((.....))))........((((((.((((((((...))))))))......((((((.......)))))).))))))..))))))........ ( -23.40) >DroEre_CAF1 2218 107 + 1 ---AGGCCUAAUAAUUUGUU-GUUUAUAGCAACUGUUUUUGAUCAGUG---CCAUUGGAUGGCAGUUUACUAUAAUUUUUAUGAUGAAUUGCUUGGCCGCGUGUGAAAUCUGCA ---.((((...(((((..(.-.(((((((..((((........))))(---((((...)))))......)))))).....)..)..)))))...))))(((.(......)))). ( -26.20) >DroYak_CAF1 2146 114 + 1 UGCAGAUUUAAUAAAUUUUUGGUUAACUGCAAGUGUUUUUGAUCAGUGCUGCCAUUGGAUGGCAGUUUACUAUAAUUUUUAUGAUGAAUUGCUUGGCCGCGUGUGAAAUCUGCA .((((((((.(((.....((((((((..((....))..)))))))).((.((((......(((((((((.((((.....)))).))))))))))))).)).))).)))))))). ( -34.70) >consensus ___AG__UUAAUAAAUUGUU_GUUUACUGCAACUGUUUUUGAUCAGUGCUGCCAUUGGAUGGCAGCUUACUAUAAUUUUUAUGAUGAAUUGCUUGGCCGCGUGUGAAAUCUGCA ............................(((((((........))))((.((((......(((((((((.((((.....)))).))))))))))))).))..........))). (-18.26 = -19.32 + 1.06)

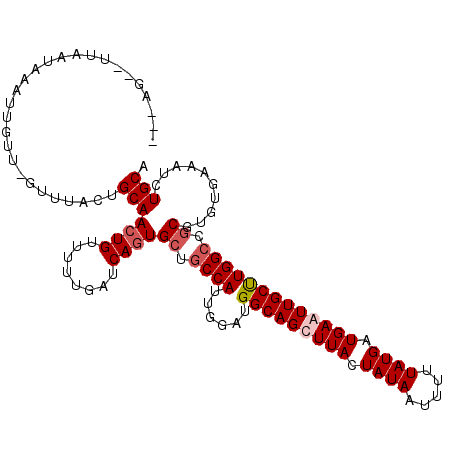

| Location | 5,583,677 – 5,583,787 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 86.50 |
| Mean single sequence MFE | -22.37 |
| Consensus MFE | -14.40 |
| Energy contribution | -15.65 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.93 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.888410 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5583677 110 - 23771897 UGCAGAUUUCACACGCGGCCAGGCAAUUCAUCAUAAAAAUUAUAGUAAGCUGCCAUCCAAUGGCAGCACUGACCAAAAACAGUUGCAGUAAGC-AACAAACUAAAAA--CUA-A ..(((.........((......))........................((((((((...)))))))).)))..........(((((.....))-)))..........--...-. ( -23.70) >DroSim_CAF1 2236 105 - 1 UGCAGAUUUCACACGCGGCCAAGCAAUUCAUCAUAAAAAUUAUAGUAAGCUGCCAUCCAAUGGCAGCACUGACCAAAAACAGUUGCAGUAAAC-AAAAAUAUAUAA-------- ((((..........((......))........................((((((((...))))))))((((........))))))))......-............-------- ( -20.60) >DroEre_CAF1 2218 107 - 1 UGCAGAUUUCACACGCGGCCAAGCAAUUCAUCAUAAAAAUUAUAGUAAACUGCCAUCCAAUGG---CACUGAUCAAAAACAGUUGCUAUAAAC-AACAAAUUAUUAGGCCU--- ................((((((....))...........(((((((((..((((((...))))---))(((........))))))))))))..-............)))).--- ( -20.90) >DroYak_CAF1 2146 114 - 1 UGCAGAUUUCACACGCGGCCAAGCAAUUCAUCAUAAAAAUUAUAGUAAACUGCCAUCCAAUGGCAGCACUGAUCAAAAACACUUGCAGUUAACCAAAAAUUUAUUAAAUCUGCA .(((((((......((......))..................((((((((((((((...))))))).((((..(((......)))))))..........)))))))))))))). ( -24.30) >consensus UGCAGAUUUCACACGCGGCCAAGCAAUUCAUCAUAAAAAUUAUAGUAAACUGCCAUCCAAUGGCAGCACUGACCAAAAACAGUUGCAGUAAAC_AAAAAAUUAUAAA__CU___ .((((.........((......))..................((((..((((((((...))))))))))))...........))))............................ (-14.40 = -15.65 + 1.25)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:22:45 2006