
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,577,917 – 5,578,019 |
| Length | 102 |
| Max. P | 0.969315 |

| Location | 5,577,917 – 5,578,019 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 83.04 |
| Mean single sequence MFE | -28.71 |
| Consensus MFE | -23.36 |
| Energy contribution | -24.03 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.60 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969315 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
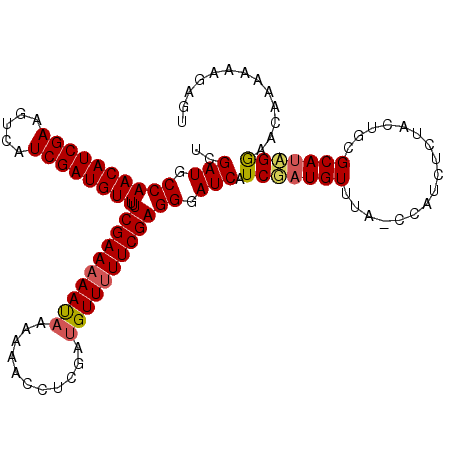
>3L_DroMel_CAF1 5577917 102 + 23771897 UCGAUGCCAACAUCGAUGUCAUCGAUGUUUUCGAAAAAUCAAAAACGUCGAUGUUUUUCGAGGAAUCGUCGAUGUUU----------UCAGCGCAUAGAAACAGCAAAGUGU ((.(((((((((((((((.((((((((((((.((....)).)))))))))))).............)))))))))).----------...).)))).))............. ( -29.54) >DroSec_CAF1 2185 112 + 1 UCGAUGCCAACAUCGAAGUCAUCGAUGUUUUCGAAAAACAAAAAACCUCCAUGUUUUUCGAGGGAUCAUCGUUGUUUAACCAUCUCUACUGCGCAUCGGAACAAAAAAGAGU ((((((((((((((((.....)))))))).((((((((((...........))))))))))(((((....(((....))).)))))....).)))))))............. ( -29.00) >DroSim_CAF1 2535 111 + 1 UCGAUGCCAACAUCGAAGUCAUCGAUGUUUUCGAAAAAUAAAAAACCUCGAUGUU-UUCGAGGGAUCAUCAAUGUUUACCCAUCUCUACUGCGCAUUGGAACAAAAAAGCGU (((((((((...((((((.((((((.(((((..........))))).)))))).)-)))))(((..((....))....)))........)).)))))))............. ( -27.60) >consensus UCGAUGCCAACAUCGAAGUCAUCGAUGUUUUCGAAAAAUAAAAAACCUCGAUGUUUUUCGAGGGAUCAUCGAUGUUUA_CCAUCUCUACUGCGCAUAGGAACAAAAAAGAGU ..(((.((((((((((.....)))))))).((((((((((...........)))))))))))).))).(((((((.................)))))))............. (-23.36 = -24.03 + 0.67)


| Location | 5,577,917 – 5,578,019 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 83.04 |
| Mean single sequence MFE | -30.13 |
| Consensus MFE | -18.89 |
| Energy contribution | -20.00 |
| Covariance contribution | 1.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.721254 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5577917 102 - 23771897 ACACUUUGCUGUUUCUAUGCGCUGA----------AAACAUCGACGAUUCCUCGAAAAACAUCGACGUUUUUGAUUUUUCGAAAACAUCGAUGACAUCGAUGUUGGCAUCGA ...................((.((.----------.(((((((((((....))).....((((((.((((((((....)))))))).))))))...))))))))..)).)). ( -31.70) >DroSec_CAF1 2185 112 - 1 ACUCUUUUUUGUUCCGAUGCGCAGUAGAGAUGGUUAAACAACGAUGAUCCCUCGAAAAACAUGGAGGUUUUUUGUUUUUCGAAAACAUCGAUGACUUCGAUGUUGGCAUCGA ..............((((((......(.(((.(((.......))).))).)((((((((((...........)))))))))).((((((((.....)))))))).)))))). ( -31.10) >DroSim_CAF1 2535 111 - 1 ACGCUUUUUUGUUCCAAUGCGCAGUAGAGAUGGGUAAACAUUGAUGAUCCCUCGAA-AACAUCGAGGUUUUUUAUUUUUCGAAAACAUCGAUGACUUCGAUGUUGGCAUCGA .((((((.((((........)))).)))).))......((((((((...((((((.-....))))))(((((........)))))))))))))...((((((....)))))) ( -27.60) >consensus ACACUUUUUUGUUCCAAUGCGCAGUAGAGAUGG_UAAACAUCGAUGAUCCCUCGAAAAACAUCGAGGUUUUUUAUUUUUCGAAAACAUCGAUGACUUCGAUGUUGGCAUCGA ......................................((((((((...((((((......))))))(((((........)))))))))))))...((((((....)))))) (-18.89 = -20.00 + 1.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:22:43 2006