
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,478,934 – 5,479,081 |
| Length | 147 |
| Max. P | 0.997548 |

| Location | 5,478,934 – 5,479,054 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.11 |
| Mean single sequence MFE | -27.29 |
| Consensus MFE | -27.10 |
| Energy contribution | -26.91 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.99 |
| SVM decision value | 2.88 |
| SVM RNA-class probability | 0.997548 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5478934 120 + 23771897 GGUGACCAUGGCAAUUAAAUCUAAAAUAUAAGCUGGGAUAUUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUG .....((.((((...................))))))..(((((((((...........)))))))))((((..(((((....((((......))))..)))))..)))).......... ( -27.01) >DroGri_CAF1 1108 120 + 1 AGUGGCCAUGGCAAUUAAAUCUAAAAUAUAAGCUGGGAUAUUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACAUACGAGAAGCACCUGUGACAAUAAUG .....((.((((...................))))))..(((((((((...........)))))))))((((..(((((.(((.........)))....)))))..)))).......... ( -26.41) >DroEre_CAF1 39669 120 + 1 AGUGACCAUGGCAAUUAAAUCUAAAAUAUAAGCCGGGAUAUUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUG .....((.((((...................))))))..(((((((((...........)))))))))((((..(((((....((((......))))..)))))..)))).......... ( -29.91) >DroYak_CAF1 28266 120 + 1 GGUGACCAUGGCAAUUAAAUCUAAAAUAUAAGCCGGGAUAUUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUG .....((.((((...................))))))..(((((((((...........)))))))))((((..(((((....((((......))))..)))))..)))).......... ( -29.91) >DroMoj_CAF1 34430 120 + 1 GGUGGCCAUGGCAAUUAAAUCUAAAAUAUAAGCUGGGAUAUUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUG ((((((....)).......(((.......(((((..(.((((((((((...........)))))))))))...))))).....((((......)))).)))..))))............. ( -27.00) >DroAna_CAF1 893 120 + 1 GGUGACCAUGGCAAUUAAAUCUAAAAUAUAAGCUGGGAUAUUGUGAAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUG .....((.((((...................)))))).((((((..........(((....)))....((((..(((((....((((......))))..)))))..)))).))))))... ( -23.51) >consensus GGUGACCAUGGCAAUUAAAUCUAAAAUAUAAGCUGGGAUAUUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUG .....((.((((...................))))))..(((((((((...........)))))))))((((..(((((....((((......))))..)))))..)))).......... (-27.10 = -26.91 + -0.19)


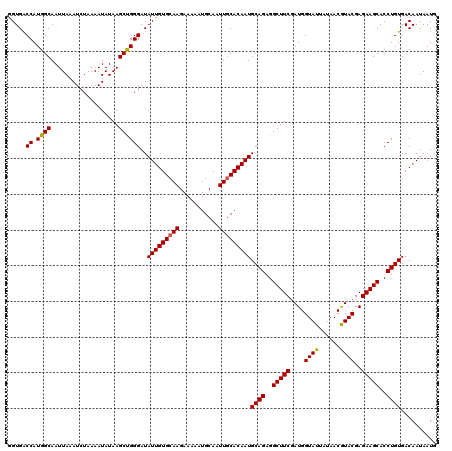
| Location | 5,478,974 – 5,479,081 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 81.72 |
| Mean single sequence MFE | -29.05 |
| Consensus MFE | -21.91 |
| Energy contribution | -21.93 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.721066 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5478974 107 + 23771897 UUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUGCGGUCGAUCGCAGGAUAUGUGGG------UGU------G ((((((((...........)))))))).((((..(((((....((((......))))..)))))..)))).(((((..(((((....)))))..)).)))...------...------. ( -29.60) >DroVir_CAF1 22380 106 + 1 UUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUGCGGUCCAAUGGGUUCGUUGUCCA------AGU------- ((((((((...........)))))))).((((..(((((....((((......))))..)))))..))))(((((((((.((......)).))).))))))..------...------- ( -30.10) >DroGri_CAF1 1148 112 + 1 UUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACAUACGAGAAGCACCUGUGACAAUAAUGCGGUCCAAUGGGUUCCGUGUCCAGUCCAAAGU------- ((((((((...........)))))))).((((..(((((.(((.........)))....)))))..))))(((....((((((.((....))..))))))...)))......------- ( -31.80) >DroWil_CAF1 3396 95 + 1 UUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUGCGGCGGAC----------------------AUACGCC-- ((((((((...........)))))))).((((..(((((....((((......))))..)))))..))))...........((((...----------------------...))))-- ( -29.30) >DroMoj_CAF1 34470 106 + 1 UUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUGUGGUCCAGCGGGUUCGAUGUCCA------UGU------- ((((((((...........)))))))).((((..(((((....((((......))))..)))))..)))).......((((((.(...((....))..).)))------)))------- ( -28.10) >DroAna_CAF1 933 107 + 1 UUGUGAAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUGCGGUCGAUCGAAAGGUGUG------------UGUGUGGG ...................(..((((..((((..(((((....((((......))))..)))))..))))(((.........))).(((....)))...------------))))..). ( -25.40) >consensus UUGUGCAAGAAAAAUGCAAUUGCACAAUGCAGAGGCUUCGAUGGUAUUAUAACGUACGAGAAGCACCUGUGACAAUAAUGCGGUCCAACGGGUUCGAUGUCCA_______GU_______ ((((((((...........)))))))).((((..(((((....((((......))))..)))))..))))................................................. (-21.91 = -21.93 + 0.03)

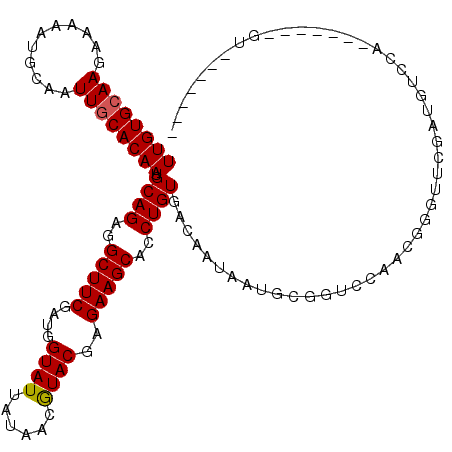

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:22:09 2006