| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,471,641 – 5,471,731 |
| Length | 90 |
| Max. P | 0.999496 |

| Location | 5,471,641 – 5,471,731 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 71.36 |
| Mean single sequence MFE | -27.25 |
| Consensus MFE | -16.96 |
| Energy contribution | -19.28 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.62 |
| SVM decision value | 3.47 |
| SVM RNA-class probability | 0.999262 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5471641 90 + 23771897 GAUGUUU--------------------------UCGUUUAUUUUUGUGGAUAGUCGGUUCCAAAUAGUUUGUUACAAACUAAAGCGGAUAGUUUGUAACGGGCUGCCUUCUUGGAU .......--------------------------(((.((((((....)))))).))).(((((.(((((((((((((((((.......))))))))))))))))).....))))). ( -25.20) >DroSec_CAF1 33185 90 + 1 GAUGUUU--------------------------UCGCUUAUUUUUGUGGAUAUUCGGUUCCAAAAAGUUUGUUACAAACUAAAGCGGUUAGUCUGUAGCGGGCUACCUUUUUGGAU ((....(--------------------------((((........)))))...))...((((((((((((((((((.(((((.....))))).)))))))))))....))))))). ( -25.90) >DroSim_CAF1 34738 90 + 1 GAUGUUU--------------------------UGGCUUAUUUUUGUGGAUAUUCGGUUCCAAACAGUUUGUUACAAACUAAAGCGUUUAGUUUGUAGCGGGCUACCGUUUUGGAU ((....(--------------------------(.((........)).))...))...((((((((((((((((((((((((.....)))))))))))))))))...).)))))). ( -25.30) >DroYak_CAF1 20639 116 + 1 GGUAUUUCGUAUAAAGUUUUUGAACAUAUGAAGACGUUUAUGGUUCAGAAAAGUCGGUUCCAAACAGGUUGUUACAAACUAAGGCAGCUAGUUUGGCGCGUGCACUUUUCUUGGAC .........((((((((((((........)))))).))))))((((((((((((((((.((((((.(((((((.........))))))).)))))).)).)).))))))).))))) ( -32.60) >consensus GAUGUUU__________________________UCGCUUAUUUUUGUGGAUAGUCGGUUCCAAACAGUUUGUUACAAACUAAAGCGGUUAGUUUGUAGCGGGCUACCUUCUUGGAU ..........................................................((((((.(((((((((((((((((.....))))))))))))))))).....)))))). (-16.96 = -19.28 + 2.31)

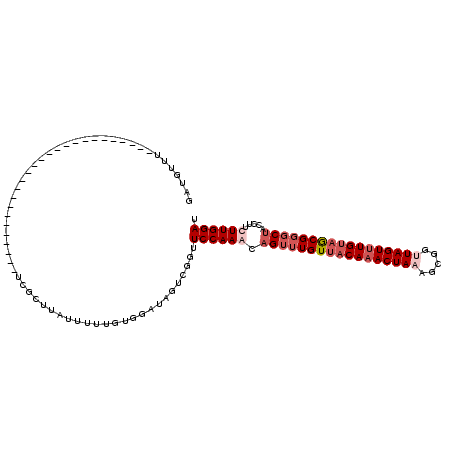

| Location | 5,471,641 – 5,471,731 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 71.36 |
| Mean single sequence MFE | -18.12 |
| Consensus MFE | -12.36 |
| Energy contribution | -13.05 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.68 |
| SVM decision value | 3.66 |
| SVM RNA-class probability | 0.999496 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
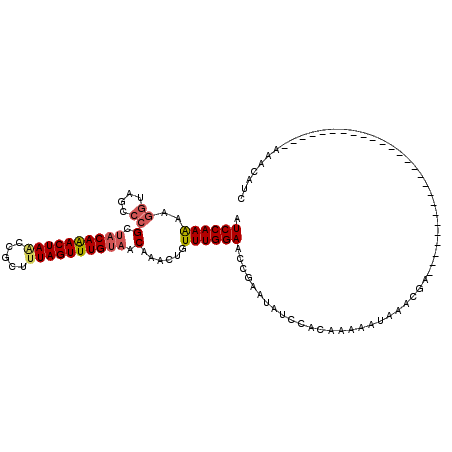
>3L_DroMel_CAF1 5471641 90 - 23771897 AUCCAAGAAGGCAGCCCGUUACAAACUAUCCGCUUUAGUUUGUAACAAACUAUUUGGAACCGACUAUCCACAAAAAUAAACGA--------------------------AAACAUC .((((((..((....))(((((((((((.......)))))))))))......)))))).........................--------------------------....... ( -19.50) >DroSec_CAF1 33185 90 - 1 AUCCAAAAAGGUAGCCCGCUACAGACUAACCGCUUUAGUUUGUAACAAACUUUUUGGAACCGAAUAUCCACAAAAAUAAGCGA--------------------------AAACAUC .(((((((((((((....))))(((((((.....)))))))........)))))))))..((..(((........)))..)).--------------------------....... ( -19.00) >DroSim_CAF1 34738 90 - 1 AUCCAAAACGGUAGCCCGCUACAAACUAAACGCUUUAGUUUGUAACAAACUGUUUGGAACCGAAUAUCCACAAAAAUAAGCCA--------------------------AAACAUC .((((((.((((.....(.((((((((((.....)))))))))).)..)))))))))).........................--------------------------....... ( -17.40) >DroYak_CAF1 20639 116 - 1 GUCCAAGAAAAGUGCACGCGCCAAACUAGCUGCCUUAGUUUGUAACAACCUGUUUGGAACCGACUUUUCUGAACCAUAAACGUCUUCAUAUGUUCAAAAACUUUAUACGAAAUACC .....((((((((.(......((((((((.....))))))))......((.....))....)))))))))((((.(((..........)))))))..................... ( -16.60) >consensus AUCCAAAAAGGUAGCCCGCUACAAACUAACCGCUUUAGUUUGUAACAAACUGUUUGGAACCGAAUAUCCACAAAAAUAAACGA__________________________AAACAUC .((((((..((....))(.((((((((((.....)))))))))).)......)))))).......................................................... (-12.36 = -13.05 + 0.69)

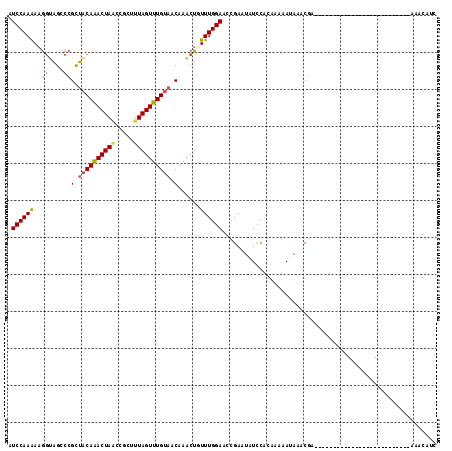
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:22:03 2006