
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,451,466 – 5,451,611 |
| Length | 145 |
| Max. P | 0.553983 |

| Location | 5,451,466 – 5,451,556 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 90.22 |
| Mean single sequence MFE | -11.47 |
| Consensus MFE | -9.98 |
| Energy contribution | -9.48 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553006 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
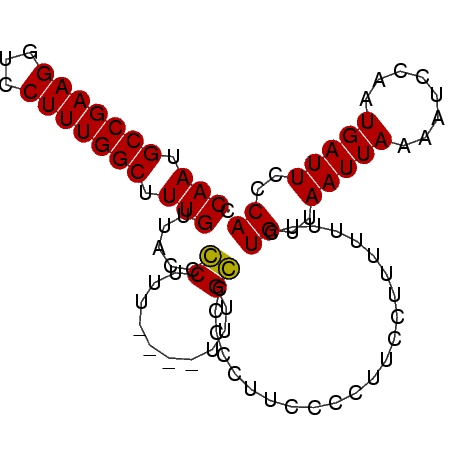
>3L_DroMel_CAF1 5451466 90 - 23771897 CCAAUGCCGAAGGUCCUUUGGCUUUGUUACCGCUUU----UUUGCCCCCUUUUCCCUUCCUUUUUUUCUGUUAAUUAAAAUCCAAUGAUUCCCA .(((.(((((((...))))))).))).....((...----...))........................(..(((((........)))))..). ( -10.70) >DroSec_CAF1 12900 94 - 1 CCAAUGCCGAAGGUCCUUUGGCUUUGUUACCCCUUUUUUUUUUGGCCCCCUUCCCCUUCCUUUUUUUUUGUUAAUUAAAAUCCAAUGAUUCCCA .(((.(((((((...))))))).))).........((((..(((((.......................)))))..)))).............. ( -10.00) >DroSim_CAF1 13027 90 - 1 CCAAUGCCGAAGGUCCUUUGGCUUUGUUACCCCUUU----UUUGGCCCCCUUCCCCUUCCUUUUUUUCUGUUAAUUAAAAUCCAAUGAUUCCCA .(((.(((((((...))))))).))).....((...----...))........................(..(((((........)))))..). ( -10.20) >DroEre_CAF1 12566 87 - 1 GCAAUGCCGAAGGUCCUUUGGCUUUGUUACCGCUUU----UUUGCUUCCUUU-CCCUU--UUUUUCGUUGUUAAUUAAAAUCCAAUGAUUCCCA ((((.(((((((...))))))).))))....((...----...)).......-.....--....((((((............))))))...... ( -15.00) >consensus CCAAUGCCGAAGGUCCUUUGGCUUUGUUACCCCUUU____UUUGCCCCCCUUCCCCUUCCUUUUUUUCUGUUAAUUAAAAUCCAAUGAUUCCCA .(((.(((((((...))))))).))).....((..........)).......................((..(((((........)))))..)) ( -9.98 = -9.48 + -0.50)


| Location | 5,451,520 – 5,451,611 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 85.27 |
| Mean single sequence MFE | -21.40 |
| Consensus MFE | -13.44 |
| Energy contribution | -14.00 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553983 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5451520 91 - 23771897 AAAAAAACACACAAAC-----------AAAGUGCAGAGAAUAUGCAUGUGUAUAUUC-------UUUUUUUUACCAAUGCCGAAGGUCCUUUGGCUUUGUUACCGCUUU--- ................-----------((((((.(((((((((((....))))))))-------))).......(((.(((((((...))))))).)))....))))))--- ( -23.60) >DroSec_CAF1 12955 106 - 1 AAA-AAACACACAAACAACGAGAGCACAAAGUGCACAGAAUAUGCAAGUGUAUCCUCC-----UUUUUUUUUACCAAUGCCGAAGGUCCUUUGGCUUUGUUACCCCUUUUUU ...-.........(((((.(((.((((....((((.......)))).))))...))).-----...............(((((((...))))))).)))))........... ( -18.00) >DroSim_CAF1 13081 103 - 1 AAA-AAACACACAAACAACGAGAGCACAAAGUGCACAGAAUAUGCAUGUGUAUCCUCC-----UUUUUUUUUACCAAUGCCGAAGGUCCUUUGGCUUUGUUACCCCUUU--- ...-.........(((((.(((.(((((...((((.......)))))))))...))).-----...............(((((((...))))))).)))))........--- ( -18.80) >DroEre_CAF1 12617 108 - 1 AAA-AAACAAACAAACAACGAGAGCACAAAGUGCACAUAAUAUGCAUGUGUAUUCUCUUCUUUUUUUUUUUUAGCAAUGCCGAAGGUCCUUUGGCUUUGUUACCGCUUU--- ...-................(((((....(((((((((.......))))))))).................((((((.(((((((...))))))).))))))..)))))--- ( -25.20) >consensus AAA_AAACACACAAACAACGAGAGCACAAAGUGCACAGAAUAUGCAUGUGUAUCCUCC_____UUUUUUUUUACCAAUGCCGAAGGUCCUUUGGCUUUGUUACCCCUUU___ ........((((......(....)......(((((.......))))))))).......................(((.(((((((...))))))).)))............. (-13.44 = -14.00 + 0.56)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:37 2006