
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,446,110 – 5,446,310 |
| Length | 200 |
| Max. P | 0.974916 |

| Location | 5,446,110 – 5,446,230 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.18 |
| Mean single sequence MFE | -27.95 |
| Consensus MFE | -21.88 |
| Energy contribution | -22.12 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553207 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5446110 120 + 23771897 UACUAUGUUGUGAGUGAUAUUCAGAGUUUAAUUUCCAUUAAGCAAAAUGCUUUUGCAUUUCGGCCGUUGGGUAAAUUUGCACUAACGCAUGUGCAUACCGAAUAUGUGCAUAUGUAUAUG ........((..(((..(((((((.(((((((....))))))).((((((....))))))......))))))).)))..)).....(((((((((((.......)))))))))))..... ( -30.20) >DroSec_CAF1 7663 118 + 1 UACUAUGUUGUGAGUGAUACUCAGAGUUUAAUUUCCAUUAAGCAAAAUGCUUUUGCAUUUCGGCCGUUGGGUAAAUUUGCACUAACACAUGUGCAUACCGAAUAUUUGCACAU--AUAUG ...(((((.(..((((.(((((((.(((((((....))))))).((((((....))))))......)))))))....(((((........))))).......))))..)))))--).... ( -29.30) >DroSim_CAF1 7703 118 + 1 UACUAUGUUGUGAGUGAUACUCAGAGUUUAAUUUCCAUUAAGCAAAAUGCUUUUGCAUUUCGGCCGUUGGGUAAAUUUGCACUAACACAUGUGCAUACCGAAUAUUUGCACAU--AUAUG ...(((((.(..((((.(((((((.(((((((....))))))).((((((....))))))......)))))))....(((((........))))).......))))..)))))--).... ( -29.30) >DroEre_CAF1 7503 106 + 1 UACUAUGUUGUGAGUGAUAUUCAGAGUUUAAUUUCCAUUAAGCAAAAUGCUUUUGCAUUCCGGCCGUUGGGCAAAUUUGCACUAACGCAUGUGCAUACCGAAU--------------ACG ((((........)))).(((((...(((((((....)))))))..........(((((....((.((((((((....))).)))))))..)))))....))))--------------).. ( -23.00) >consensus UACUAUGUUGUGAGUGAUACUCAGAGUUUAAUUUCCAUUAAGCAAAAUGCUUUUGCAUUUCGGCCGUUGGGUAAAUUUGCACUAACACAUGUGCAUACCGAAUAUUUGCACAU__AUAUG ......(((((((((...)))))..(((((((....))))))).((((((....)))))))))).(((.((((....(((((........))))))))).)))................. (-21.88 = -22.12 + 0.25)


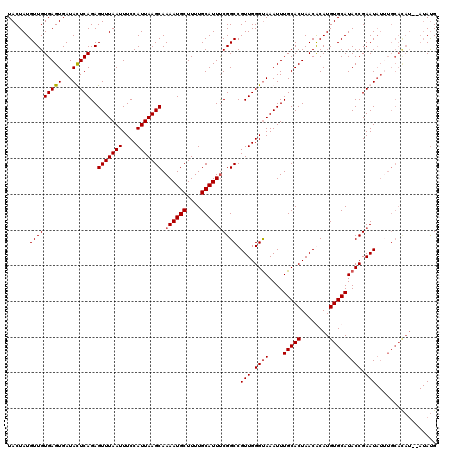
| Location | 5,446,110 – 5,446,230 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.18 |
| Mean single sequence MFE | -26.38 |
| Consensus MFE | -23.64 |
| Energy contribution | -23.70 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.74 |
| SVM RNA-class probability | 0.974916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5446110 120 - 23771897 CAUAUACAUAUGCACAUAUUCGGUAUGCACAUGCGUUAGUGCAAAUUUACCCAACGGCCGAAAUGCAAAAGCAUUUUGCUUAAUGGAAAUUAAACUCUGAAUAUCACUCACAACAUAGUA ...............((((((((..(((((........)))))(((((.((....(((.(((((((....))))))))))....))))))).....))))))))................ ( -24.50) >DroSec_CAF1 7663 118 - 1 CAUAU--AUGUGCAAAUAUUCGGUAUGCACAUGUGUUAGUGCAAAUUUACCCAACGGCCGAAAUGCAAAAGCAUUUUGCUUAAUGGAAAUUAAACUCUGAGUAUCACUCACAACAUAGUA .((((--(((((((.((.....)).))))))))))).(((...(((((.((....(((.(((((((....))))))))))....)))))))..))).((((.....)))).......... ( -28.30) >DroSim_CAF1 7703 118 - 1 CAUAU--AUGUGCAAAUAUUCGGUAUGCACAUGUGUUAGUGCAAAUUUACCCAACGGCCGAAAUGCAAAAGCAUUUUGCUUAAUGGAAAUUAAACUCUGAGUAUCACUCACAACAUAGUA .((((--(((((((.((.....)).))))))))))).(((...(((((.((....(((.(((((((....))))))))))....)))))))..))).((((.....)))).......... ( -28.30) >DroEre_CAF1 7503 106 - 1 CGU--------------AUUCGGUAUGCACAUGCGUUAGUGCAAAUUUGCCCAACGGCCGGAAUGCAAAAGCAUUUUGCUUAAUGGAAAUUAAACUCUGAAUAUCACUCACAACAUAGUA .((--------------((((((..(((((........)))))(((((.((....(((.(((((((....))))))))))....))))))).....))))))))................ ( -24.40) >consensus CAUAU__AUGUGCAAAUAUUCGGUAUGCACAUGCGUUAGUGCAAAUUUACCCAACGGCCGAAAUGCAAAAGCAUUUUGCUUAAUGGAAAUUAAACUCUGAAUAUCACUCACAACAUAGUA ...............((((((((..(((((........)))))(((((.((....(((.(((((((....))))))))))....))))))).....))))))))................ (-23.64 = -23.70 + 0.06)


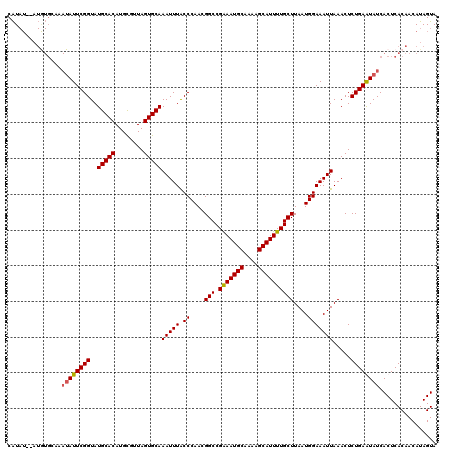
| Location | 5,446,150 – 5,446,270 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.90 |
| Mean single sequence MFE | -29.82 |
| Consensus MFE | -23.70 |
| Energy contribution | -25.57 |
| Covariance contribution | 1.87 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.44 |
| SVM RNA-class probability | 0.953944 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5446150 120 + 23771897 AGCAAAAUGCUUUUGCAUUUCGGCCGUUGGGUAAAUUUGCACUAACGCAUGUGCAUACCGAAUAUGUGCAUAUGUAUAUGGCCAUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUU .((.((((((....)))))).((((((((((((....))).)))))(((((((((((.......)))))))))))....))))..........))......................... ( -34.90) >DroSec_CAF1 7703 118 + 1 AGCAAAAUGCUUUUGCAUUUCGGCCGUUGGGUAAAUUUGCACUAACACAUGUGCAUACCGAAUAUUUGCACAU--AUAUGGUCAUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUU .((.((((((....))))))..))((((((((.....(((((........)))))..((((((((.((..((.--...))..)).))))))))...............)))))))).... ( -29.10) >DroSim_CAF1 7743 118 + 1 AGCAAAAUGCUUUUGCAUUUCGGCCGUUGGGUAAAUUUGCACUAACACAUGUGCAUACCGAAUAUUUGCACAU--AUAUGGCCAUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUU .((.((((((....)))))).((((((((((((....))).)))))..(((((((...........)))))))--....))))..........))......................... ( -30.10) >DroEre_CAF1 7543 106 + 1 AGCAAAAUGCUUUUGCAUUCCGGCCGUUGGGCAAAUUUGCACUAACGCAUGUGCAUACCGAAU--------------ACGGCCUUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUU .((..(((((....)))))..(((((((((.......(((((........)))))..)))...--------------))))))..........))......................... ( -25.20) >consensus AGCAAAAUGCUUUUGCAUUUCGGCCGUUGGGUAAAUUUGCACUAACACAUGUGCAUACCGAAUAUUUGCACAU__AUAUGGCCAUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUU .((..(((((....)))))...))((((((((.....(((((........)))))..((((((((.((..(((....)))..)).))))))))...............)))))))).... (-23.70 = -25.57 + 1.87)



| Location | 5,446,150 – 5,446,270 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.90 |
| Mean single sequence MFE | -26.17 |
| Consensus MFE | -20.98 |
| Energy contribution | -21.35 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.884195 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5446150 120 - 23771897 AAUUCAUUGAAUUUAAUAAUUUUUUGCCCAAAUAUAUGGCCAUAUACAUAUGCACAUAUUCGGUAUGCACAUGCGUUAGUGCAAAUUUACCCAACGGCCGAAAUGCAAAAGCAUUUUGCU .........................(((.....(((((..((((....))))..)))))..(((((((((........)))))....))))....)))((((((((....)))))))).. ( -25.10) >DroSec_CAF1 7703 118 - 1 AAUUCAUUGAAUUUAAUAAUUUUUUGCCCAAAUAUAUGACCAUAU--AUGUGCAAAUAUUCGGUAUGCACAUGUGUUAGUGCAAAUUUACCCAACGGCCGAAAUGCAAAAGCAUUUUGCU ........((((..........(((((....((((((....))))--))..))))).))))(((((((((........)))))....))))....(((.(((((((....)))))))))) ( -24.20) >DroSim_CAF1 7743 118 - 1 AAUUCAUUGAAUUUAAUAAUUUUUUGCCCAAAUAUAUGGCCAUAU--AUGUGCAAAUAUUCGGUAUGCACAUGUGUUAGUGCAAAUUUACCCAACGGCCGAAAUGCAAAAGCAUUUUGCU ....((..(((((....)))))..))...........((((((((--(((((((.((.....)).)))))))))))..(((......))).....))))(((((((....)))))))... ( -27.80) >DroEre_CAF1 7543 106 - 1 AAUUCAUUGAAUUUAAUAAUUUUUUGCCCAAAUAUAAGGCCGU--------------AUUCGGUAUGCACAUGCGUUAGUGCAAAUUUGCCCAACGGCCGGAAUGCAAAAGCAUUUUGCU ....((..(((((....)))))..))...........((((((--------------....(((((((((........)))))....))))..))))))(((((((....)))))))... ( -27.60) >consensus AAUUCAUUGAAUUUAAUAAUUUUUUGCCCAAAUAUAUGGCCAUAU__AUGUGCAAAUAUUCGGUAUGCACAUGCGUUAGUGCAAAUUUACCCAACGGCCGAAAUGCAAAAGCAUUUUGCU ....((..(((((....)))))..))...........((((......((((....))))..(((((((((........)))))....))))....))))(((((((....)))))))... (-20.98 = -21.35 + 0.37)

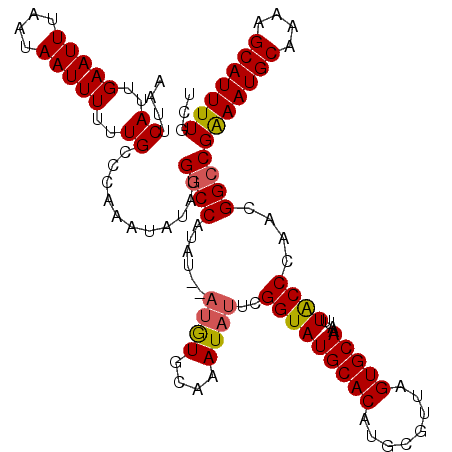
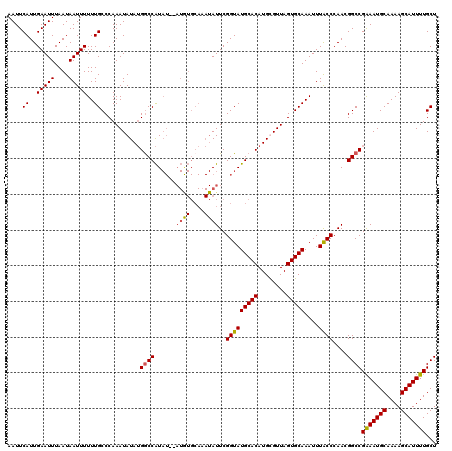
| Location | 5,446,190 – 5,446,310 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.32 |
| Mean single sequence MFE | -24.98 |
| Consensus MFE | -19.54 |
| Energy contribution | -20.97 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.686420 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5446190 120 + 23771897 ACUAACGCAUGUGCAUACCGAAUAUGUGCAUAUGUAUAUGGCCAUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUUAUUUAUUUAGCAGGUGGCGCUGACUGAAUAAACCCGUAUA ......(((((((((((.......)))))))))))(((((((((......)))............................((((((((((((......))).))))))))).)))))). ( -30.00) >DroSec_CAF1 7743 118 + 1 ACUAACACAUGUGCAUACCGAAUAUUUGCACAU--AUAUGGUCAUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUUAUUUAUUUAGCAGGUGGCGCUGACUGAAUAAACCCGCAUA .........(((((...((((((((.((..((.--...))..)).))))))))............................((((((((((((......))).)))))))))...))))) ( -24.00) >DroSim_CAF1 7783 118 + 1 ACUAACACAUGUGCAUACCGAAUAUUUGCACAU--AUAUGGCCAUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUUAUUUAUUUAGCAGGUGGCGCUGACUGAAUAAACCCGCAUA ........(((((((...........)))))))--...((.(((......))).)).................(((.....((((((((((((......))).)))))))))....))). ( -24.40) >DroEre_CAF1 7583 106 + 1 ACUAACGCAUGUGCAUACCGAAU--------------ACGGCCUUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUUAUUUAUUUAGCAGGUGGCGCUGACUGAAUAAACCCGCAUA .........(((((...(((...--------------.)))..........(((.............(((((...))))).((((((((((((......))).))))))))))))))))) ( -21.50) >consensus ACUAACACAUGUGCAUACCGAAUAUUUGCACAU__AUAUGGCCAUAUAUUUGGGCAAAAAAUUAUUAAAUUCAAUGAAUUAUUUAUUUAGCAGGUGGCGCUGACUGAAUAAACCCGCAUA .........(((((...((((((((.((..(((....)))..)).))))))))............................((((((((((((......))).)))))))))...))))) (-19.54 = -20.97 + 1.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:22 2006