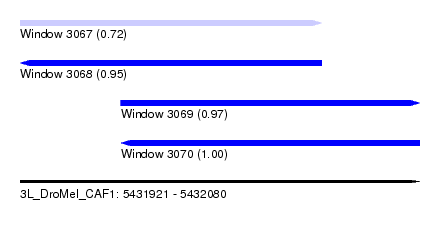

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,431,921 – 5,432,080 |
| Length | 159 |
| Max. P | 0.998485 |

| Location | 5,431,921 – 5,432,041 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -42.75 |
| Consensus MFE | -35.08 |
| Energy contribution | -36.20 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.719228 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5431921 120 + 23771897 AUAGGGGAGUCGUUUCUCGUUCCACCUGCUAAUUCGAAUUUAAUGCCUGCUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCAAAAGGUGAGCAGGAAGGGGA .((((((((.((.....)))))).))))...............(.(((.(((((((.(..((((.((((.....))))..))))(((((......)))))...).)))))))..))).). ( -37.90) >DroSec_CAF1 2092 120 + 1 GUGGGGGAGUCGUUUCUCGUUCCACCUGCUAAUUCGAAUUUAAUGCCUGCUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCGACAGGUGAGCAGGAAGCGUA (..((((((.((.....)))))).))..).............((((.(.(((((((.((((.....))))........((((((.((((......))))))))))))))))).).)))). ( -46.90) >DroSim_CAF1 2166 120 + 1 AUGGGGGAGUCCUUUCUCGUUCCACCUGCUAAUUCGAAUUUAAUGCCUGCUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCGACAGGUGAGCAGGAAGCGGA .((((((((......))).)))))...................(((.(.(((((((.((((.....))))........((((((.((((......))))))))))))))))).).))).. ( -40.50) >DroEre_CAF1 2016 119 + 1 ACGGGGGACACCCUUCUCGUUCCACCUGCUAAUUCGAAUUUAAUGCCU-CUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCGACAGGUGAGCAGGAGGCGGA ..(((((((.........))))).)).................(((((-(((((((.((((.....))))........((((((.((((......)))))))))))))))).)))))).. ( -45.70) >consensus AUGGGGGAGUCCUUUCUCGUUCCACCUGCUAAUUCGAAUUUAAUGCCUGCUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCGACAGGUGAGCAGGAAGCGGA .((((((((.((.....)))))).))))...............(((.(.(((((((.((((.....))))........((((((.((((......))))))))))))))))).).))).. (-35.08 = -36.20 + 1.12)


| Location | 5,431,921 – 5,432,041 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -39.32 |
| Consensus MFE | -36.06 |
| Energy contribution | -36.00 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.947010 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5431921 120 - 23771897 UCCCCUUCCUGCUCACCUUUUGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAGCAGGCAUUAAAUUCGAAUUAGCAGGUGGAACGAGAAACGACUCCCCUAU ...((..((((((..((((((((((......))))))).))).....((((.((((((......))))))...)))).............)))))).))...(((......)))...... ( -33.50) >DroSec_CAF1 2092 120 - 1 UACGCUUCCUGCUCACCUGUCGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAGCAGGCAUUAAAUUCGAAUUAGCAGGUGGAACGAGAAACGACUCCCCCAC ...((((.(((((((((((((((((......)))).))))))........((((.....)))).))))))).))))...................((((...(((......)))..)))) ( -40.40) >DroSim_CAF1 2166 120 - 1 UCCGCUUCCUGCUCACCUGUCGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAGCAGGCAUUAAAUUCGAAUUAGCAGGUGGAACGAGAAAGGACUCCCCCAU ...((((.(((((((((((((((((......)))).))))))........((((.....)))).))))))).))))..................((.((...(((......))))))).. ( -39.90) >DroEre_CAF1 2016 119 - 1 UCCGCCUCCUGCUCACCUGUCGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAG-AGGCAUUAAAUUCGAAUUAGCAGGUGGAACGAGAAGGGUGUCCCCCGU (((((((.(((((((((((((((((......)))).))))))........((((.....)))).)))))))-..((...............))))))))).......(((.....))).. ( -43.46) >consensus UCCGCUUCCUGCUCACCUGUCGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAGCAGGCAUUAAAUUCGAAUUAGCAGGUGGAACGAGAAACGACUCCCCCAU ...((((.(((((((((((((((((......)))).))))))........((((.....)))).))))))).))))..................((.(((............))).)).. (-36.06 = -36.00 + -0.06)


| Location | 5,431,961 – 5,432,080 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 87.16 |
| Mean single sequence MFE | -40.15 |
| Consensus MFE | -33.70 |
| Energy contribution | -34.70 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.68 |
| SVM RNA-class probability | 0.971663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5431961 119 + 23771897 UAAUGCCUGCUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCAAAAGGUGAGCAGGAAGGGGAGGAGGGCGGCUUGUUUGCGGUUGUUCACGCCAGAAGGGC ....((((.(((((((.(..((((.((((.....))))..))))(((((......)))))...).)))))))...((.(.(((.(((.((......)).))).))).).))....)))) ( -43.30) >DroSec_CAF1 2132 99 + 1 UAAUGCCUGCUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCGACAGGUGAGCAGGAAGCGUAGGAGUGCGACUUGUUUGCG-------------------- ..((((.(.(((((((.((((.....))))........((((((.((((......))))))))))))))))).).))))......((((.....)))).-------------------- ( -37.60) >DroSim_CAF1 2206 99 + 1 UAAUGCCUGCUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCGACAGGUGAGCAGGAAGCGGAGGAGUGCGGCUUGUUUGCG-------------------- .....(((((((((...((((.....))))...)))..((((((.((((......))))))))))..))))))..(((((.(((.....))).))))).-------------------- ( -39.30) >DroEre_CAF1 2056 98 + 1 UAAUGCCU-CUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCGACAGGUGAGCAGGAGGCGGAGGAGUGCGGCUUGUUUGCG-------------------- ....((((-(((((((.((((.....))))........((((((.((((......)))))))))))))))).)))))(((.(((.....))).)))...-------------------- ( -40.40) >consensus UAAUGCCUGCUGCUCAACAUGACAAACAUGGCAAGUGUCCUGUCUGCCAGGUGAAUGGCGACAGGUGAGCAGGAAGCGGAGGAGUGCGGCUUGUUUGCG____________________ .........(((((((.((((.....))))........((((((.((((......)))))))))))))))))...(((((.(((.....))).)))))..................... (-33.70 = -34.70 + 1.00)


| Location | 5,431,961 – 5,432,080 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 87.16 |
| Mean single sequence MFE | -35.83 |
| Consensus MFE | -32.65 |
| Energy contribution | -32.77 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.57 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.12 |
| SVM RNA-class probability | 0.998485 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5431961 119 - 23771897 GCCCUUCUGGCGUGAACAACCGCAAACAAGCCGCCCUCCUCCCCUUCCUGCUCACCUUUUGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAGCAGGCAUUA (((.....((((....)....((......)).)))............(((((((((((((((((......))))))).)))........((((.....)))).)))))))..))).... ( -34.50) >DroSec_CAF1 2132 99 - 1 --------------------CGCAAACAAGUCGCACUCCUACGCUUCCUGCUCACCUGUCGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAGCAGGCAUUA --------------------.((.........))........((((.(((((((((((((((((......)))).))))))........((((.....)))).))))))).)))).... ( -35.40) >DroSim_CAF1 2206 99 - 1 --------------------CGCAAACAAGCCGCACUCCUCCGCUUCCUGCUCACCUGUCGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAGCAGGCAUUA --------------------.((......))...........((((.(((((((((((((((((......)))).))))))........((((.....)))).))))))).)))).... ( -35.50) >DroEre_CAF1 2056 98 - 1 --------------------CGCAAACAAGCCGCACUCCUCCGCCUCCUGCUCACCUGUCGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAG-AGGCAUUA --------------------.((......))...........(((((.((((((((((((((((......)))).))))))........((((.....)))).)))))))-)))).... ( -37.90) >consensus ____________________CGCAAACAAGCCGCACUCCUCCGCUUCCUGCUCACCUGUCGCCAUUCACCUGGCAGACAGGACACUUGCCAUGUUUGUCAUGUUGAGCAGCAGGCAUUA .....................((.........))........((((.(((((((((((((((((......)))).))))))........((((.....)))).))))))).)))).... (-32.65 = -32.77 + 0.12)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:10 2006