
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,414,702 – 5,414,812 |
| Length | 110 |
| Max. P | 0.931604 |

| Location | 5,414,702 – 5,414,812 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 94.19 |
| Mean single sequence MFE | -39.84 |
| Consensus MFE | -28.72 |
| Energy contribution | -28.76 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.29 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.826154 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
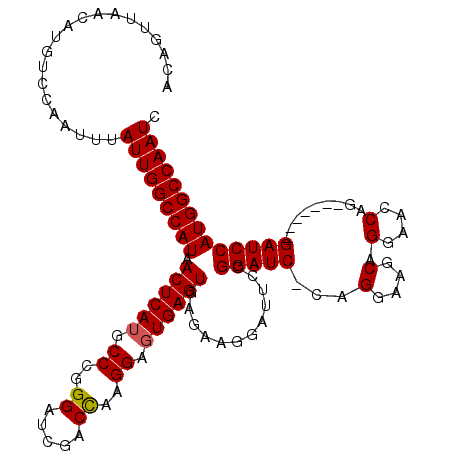
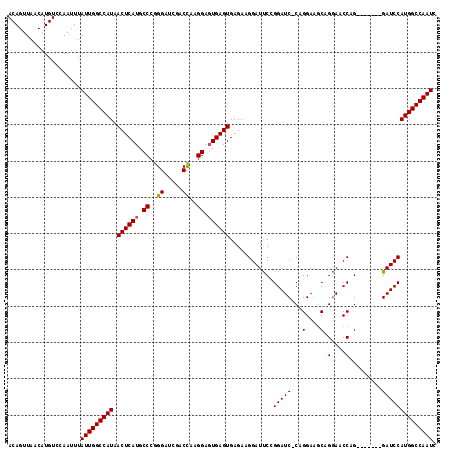
>3L_DroMel_CAF1 5414702 110 + 23771897 ACAGUUAACAUGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGACUGAGUGAGAAGGAUUCCGGAUC-CAGGAAGCAGAAACCAG-------GAUCCAUGGCCAAUC ....................(((((((((.(((((..((..((.....))..))..)))))............(((((-(.((.........)).)-------)))))))))))))). ( -34.20) >DroSec_CAF1 10288 111 + 1 ACAGUUAACAUGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGAGUGAGUGAGAAGGAUUCCGGAUCCCAGGAAGCAGGAACCAG-------GAUCCAUGGCCAAUC ....................(((((((((.((((((.((..((.....))..)).)))))).....((((((.((.(((..(....).))).)).)-------)))))))))))))). ( -38.70) >DroSim_CAF1 11479 111 + 1 ACAGUUAACAUGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGAGUGAGUGAGAAGGAUUCCGGAUCCCAGGAAGCAGGAACCAG-------GAUCCAUGGCCAAUC ....................(((((((((.((((((.((..((.....))..)).)))))).....((((((.((.(((..(....).))).)).)-------)))))))))))))). ( -38.70) >DroEre_CAF1 10192 110 + 1 ACAGUUAACAUGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGAGUGAGUGAGAAGGAUCCAGGAUC-CAGGAAGCAGGAACCAG-------GAUCCAUGGCCAAUC ....................(((((((((.((((((.((..((.....))..)).)))))).....((((((.((.((-(.(....).))).)).)-------)))))))))))))). ( -41.90) >DroYak_CAF1 9844 117 + 1 ACAGUUAACAUGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACUGAGGAGUGAGUGAGAAGGAUCCAGGAUC-CAGGAACCAGGAACCAGGAUUCAGGAUCCAUGGCCAAUC ....................(((((((((.((((((.(((((......))).)).)))))).....((((((.(((((-(.((.........)).)))))).))))))))))))))). ( -45.70) >consensus ACAGUUAACAUGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGAGUGAGUGAGAAGGAUUCCGGAUC_CAGGAAGCAGGAACCAG_______GAUCCAUGGCCAAUC ....................(((((((((.((((((.((..((.....))..)).))))))............(((((...(....).(....).........)))))))))))))). (-28.72 = -28.76 + 0.04)

| Location | 5,414,702 – 5,414,812 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 94.19 |
| Mean single sequence MFE | -39.04 |
| Consensus MFE | -29.68 |
| Energy contribution | -30.52 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.931604 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
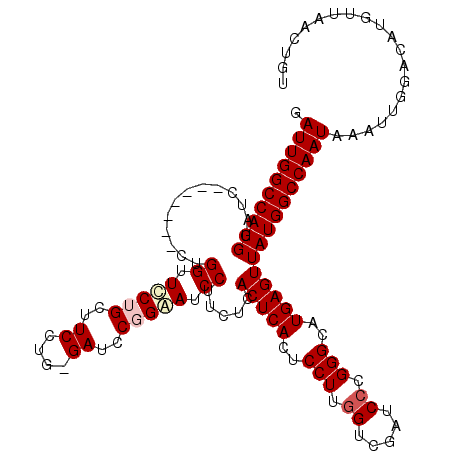
>3L_DroMel_CAF1 5414702 110 - 23771897 GAUUGGCCAUGGAUC-------CUGGUUUCUGCUUCCUG-GAUCCGGAAUCCUUCUCACUCAGUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACAUGUUAACUGU .((((((((((((((-------(.((.........)).)-)))))............(((((((((..((.....)).)))).))))).))))))))).................... ( -37.20) >DroSec_CAF1 10288 111 - 1 GAUUGGCCAUGGAUC-------CUGGUUCCUGCUUCCUGGGAUCCGGAAUCCUUCUCACUCACUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACAUGUUAACUGU .(((((((((((((.-------((((.((((.......)))).)))).)))).....(((((..(((.((.....)).)))..))))).))))))))).................... ( -39.50) >DroSim_CAF1 11479 111 - 1 GAUUGGCCAUGGAUC-------CUGGUUCCUGCUUCCUGGGAUCCGGAAUCCUUCUCACUCACUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACAUGUUAACUGU .(((((((((((((.-------((((.((((.......)))).)))).)))).....(((((..(((.((.....)).)))..))))).))))))))).................... ( -39.50) >DroEre_CAF1 10192 110 - 1 GAUUGGCCAUGGAUC-------CUGGUUCCUGCUUCCUG-GAUCCUGGAUCCUUCUCACUCACUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACAUGUUAACUGU .((((((((((((((-------(.((.(((........)-)).)).)))))).....(((((..(((.((.....)).)))..))))).))))))))).................... ( -41.90) >DroYak_CAF1 9844 117 - 1 GAUUGGCCAUGGAUCCUGAAUCCUGGUUCCUGGUUCCUG-GAUCCUGGAUCCUUCUCACUCACUCCUCAGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACAUGUUAACUGU .(((((((((((((((.(.((((.((.........)).)-))).).)))))).....(((((..(((..(.....)..)))..))))).))))))))).................... ( -37.10) >consensus GAUUGGCCAUGGAUC_______CUGGUUCCUGCUUCCUG_GAUCCGGAAUCCUUCUCACUCACUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACAUGUUAACUGU .((((((((((.............((.(((((..((....))..))))).)).....(((((..(((.((.....)).)))..))))))))))))))).................... (-29.68 = -30.52 + 0.84)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:03 2006