
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,400,024 – 5,400,144 |
| Length | 120 |
| Max. P | 0.851165 |

| Location | 5,400,024 – 5,400,144 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.39 |
| Mean single sequence MFE | -40.97 |
| Consensus MFE | -33.53 |
| Energy contribution | -33.20 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.765257 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5400024 120 + 23771897 AUCCACAUUCAGGCAUCUGCAAACAAUUCCAUGUGCGCUGGCUGUCAGAUCUGCAGGAAUUGCUCAGCUGCCUGGCCGACGAUUCUCCUGUCGCGGAAAGAGACAGCCUGAAUGGGCAGA ..((.(((((((((.(((((...(((((((.((((..(((.....)))...)))))))))))....))...((..(((.((((......)))))))..)))))..))))))))))).... ( -42.10) >DroSec_CAF1 48890 104 + 1 AUCCACAUUCAGGCAUCUGCAAACAAUUCCAUUUGCGCUGGCUGUCAGAUCUGCAGGAAUUGCUCAGCUGCCUGGCCGACGAUUCUCC----------------AGCCUGAAUGGGCAGA ..((.(((((((((....(((((........)))))..((((((.((....)))))((((((.((.(((....))).)))))))).))----------------)))))))))))).... ( -35.50) >DroYak_CAF1 48865 120 + 1 AUCCACAUUCAGGCGCCUGCUAACAAUUCCAUUUGCGCUGGCUGUCAGAUCUGCAGGAAUUGCUCAGCUGCCUGGUCGACAAUUCUCCUGUCGCAGAAAGAGACAGCCUGAGUGGGGAGA .(((.((((((((((((.((...(((......))).)).)))((((...(((((((((((((.((.(((....))).))))))..))))...)))))....))))))))))))).))).. ( -45.30) >consensus AUCCACAUUCAGGCAUCUGCAAACAAUUCCAUUUGCGCUGGCUGUCAGAUCUGCAGGAAUUGCUCAGCUGCCUGGCCGACGAUUCUCCUGUCGC_GAAAGAGACAGCCUGAAUGGGCAGA ..((.(((((((((..((((...(((......)))..(((.....)))....))))((((((.((.(((....))).))))))))....................))))))))))).... (-33.53 = -33.20 + -0.33)

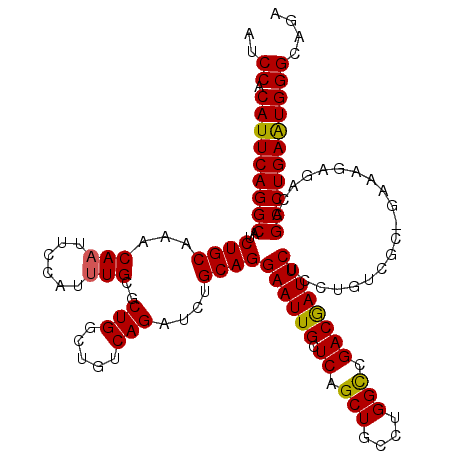

| Location | 5,400,024 – 5,400,144 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.39 |
| Mean single sequence MFE | -43.40 |
| Consensus MFE | -35.10 |
| Energy contribution | -35.77 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.851165 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5400024 120 - 23771897 UCUGCCCAUUCAGGCUGUCUCUUUCCGCGACAGGAGAAUCGUCGGCCAGGCAGCUGAGCAAUUCCUGCAGAUCUGACAGCCAGCGCACAUGGAAUUGUUUGCAGAUGCCUGAAUGUGGAU ....(((((((.(((((.(..(((((......)))))...).)))))(((((.(((.(((((((((((....(((.....))).)))...))))))))...))).)))))))))).)).. ( -45.90) >DroSec_CAF1 48890 104 - 1 UCUGCCCAUUCAGGCU----------------GGAGAAUCGUCGGCCAGGCAGCUGAGCAAUUCCUGCAGAUCUGACAGCCAGCGCAAAUGGAAUUGUUUGCAGAUGCCUGAAUGUGGAU ....(((((((.((((----------------((.......))))))(((((.(((.(((((((((((....(((.....))).)))...))))))))...))).)))))))))).)).. ( -39.70) >DroYak_CAF1 48865 120 - 1 UCUCCCCACUCAGGCUGUCUCUUUCUGCGACAGGAGAAUUGUCGACCAGGCAGCUGAGCAAUUCCUGCAGAUCUGACAGCCAGCGCAAAUGGAAUUGUUAGCAGGCGCCUGAAUGUGGAU ..(((.((.((((((.((((...((((((((((.....))))))..))))..(((((.((((((((((....(((.....))).)))...)))))))))))))))))))))).)).))). ( -44.60) >consensus UCUGCCCAUUCAGGCUGUCUCUUUC_GCGACAGGAGAAUCGUCGGCCAGGCAGCUGAGCAAUUCCUGCAGAUCUGACAGCCAGCGCAAAUGGAAUUGUUUGCAGAUGCCUGAAUGUGGAU ....(((((((((((.........................(((.....))).((.(((((((((((((....(((.....))).)))...))))))))))))....))))))))).)).. (-35.10 = -35.77 + 0.67)


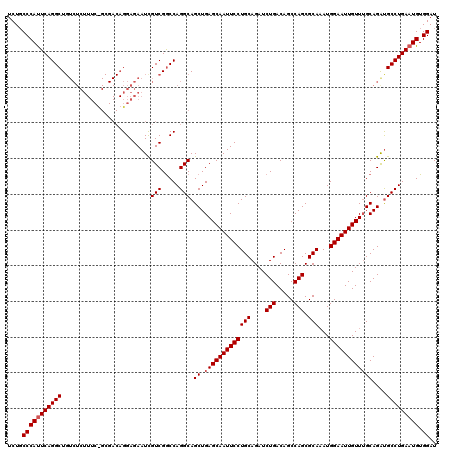
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:20:44 2006