
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,395,463 – 5,395,566 |
| Length | 103 |
| Max. P | 0.985384 |

| Location | 5,395,463 – 5,395,566 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 94.93 |
| Mean single sequence MFE | -26.45 |
| Consensus MFE | -23.15 |
| Energy contribution | -23.40 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.790108 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5395463 103 + 23771897 GACAUUGACCCUUUGCAUGUCCAAAGGGACAAAAGGGUGUUACUGUGUCACAGACGUAAUUAAUUGUAAAUAAUUGUGCAUGUGUCACUUGCAAUUUAUUUAU ((((((((((((((...(((((....))))).))))).))))..)))))................((((((((...((((.(.....).))))..)))))))) ( -28.10) >DroSec_CAF1 44323 101 + 1 GACUUUGACCCUUUGCACGUCCG-AGGGACA-AAGGGUGUUACUGUGUCACAGACGUAAUUAAUUGUAAAUAAUUGUGCAUGUGUCACUUGCAAUUUAUUUAU .......((((((((..(.....-..)..))-))))))((....(((.((((..((((((((........))))))))..)))).)))..))........... ( -24.60) >DroSim_CAF1 44134 101 + 1 GACAUUGACCCUUUGCACGUCCG-AGGGACA-AAGGGUGUUACUGUGUCACAGACGUAAUUAAUUGUAAAUAAUUGUGCGUGUGUCACUUGCAAUUUAUUUAU ((((((((((((((....((((.-..)))).-))))).))))..)))))(((.(((((..((((((....))))))))))).))).................. ( -24.70) >DroYak_CAF1 44115 101 + 1 GACAUUGACCCUUGGCACCUCCA-AGGGACA-AAGGGUGUUACUGUGUCACAGACGUAAUUAAUUGUAAAUAAUUGUGCAUGUGUCACUUGUAAUUUUUUUAU .....((.(((((((.....)))-)))).))-(((((.(((((.(((.((((..((((((((........))))))))..)))).)))..))))).))))).. ( -28.40) >consensus GACAUUGACCCUUUGCACGUCCA_AGGGACA_AAGGGUGUUACUGUGUCACAGACGUAAUUAAUUGUAAAUAAUUGUGCAUGUGUCACUUGCAAUUUAUUUAU .......((((((.....((((....))))..))))))......(((.((((..((((((((........))))))))..)))).)))............... (-23.15 = -23.40 + 0.25)

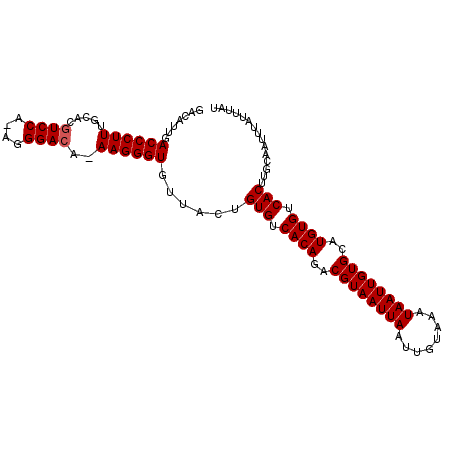

| Location | 5,395,463 – 5,395,566 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 94.93 |
| Mean single sequence MFE | -23.40 |
| Consensus MFE | -22.48 |
| Energy contribution | -22.35 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.96 |
| SVM decision value | 2.01 |
| SVM RNA-class probability | 0.985384 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5395463 103 - 23771897 AUAAAUAAAUUGCAAGUGACACAUGCACAAUUAUUUACAAUUAAUUACGUCUGUGACACAGUAACACCCUUUUGUCCCUUUGGACAUGCAAAGGGUCAAUGUC ........((((...(((.((((.((..(((((........)))))..)).)))).)))......((((((((((((....)))))...)))))))))))... ( -24.00) >DroSec_CAF1 44323 101 - 1 AUAAAUAAAUUGCAAGUGACACAUGCACAAUUAUUUACAAUUAAUUACGUCUGUGACACAGUAACACCCUU-UGUCCCU-CGGACGUGCAAAGGGUCAAAGUC ................((((...((((((((((........)))))..(((((.((((.((.......)).-))))...-))))))))))....))))..... ( -22.60) >DroSim_CAF1 44134 101 - 1 AUAAAUAAAUUGCAAGUGACACACGCACAAUUAUUUACAAUUAAUUACGUCUGUGACACAGUAACACCCUU-UGUCCCU-CGGACGUGCAAAGGGUCAAUGUC ........((((...(((.((((.((..(((((........)))))..)).)))).)))......((((((-((..(..-.....)..))))))))))))... ( -22.00) >DroYak_CAF1 44115 101 - 1 AUAAAAAAAUUACAAGUGACACAUGCACAAUUAUUUACAAUUAAUUACGUCUGUGACACAGUAACACCCUU-UGUCCCU-UGGAGGUGCCAAGGGUCAAUGUC .........((((..(((.((((.((..(((((........)))))..)).)))).))).))))......(-((.((((-(((.....))))))).))).... ( -25.00) >consensus AUAAAUAAAUUGCAAGUGACACAUGCACAAUUAUUUACAAUUAAUUACGUCUGUGACACAGUAACACCCUU_UGUCCCU_CGGACGUGCAAAGGGUCAAUGUC .........((((..(((.((((.((..(((((........)))))..)).)))).))).)))).((((((.(((((....)))))....))))))....... (-22.48 = -22.35 + -0.13)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:20:37 2006