
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,385,302 – 5,385,551 |
| Length | 249 |
| Max. P | 0.826700 |

| Location | 5,385,302 – 5,385,393 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 93.16 |
| Mean single sequence MFE | -28.70 |
| Consensus MFE | -21.28 |
| Energy contribution | -21.28 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.46 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.826700 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
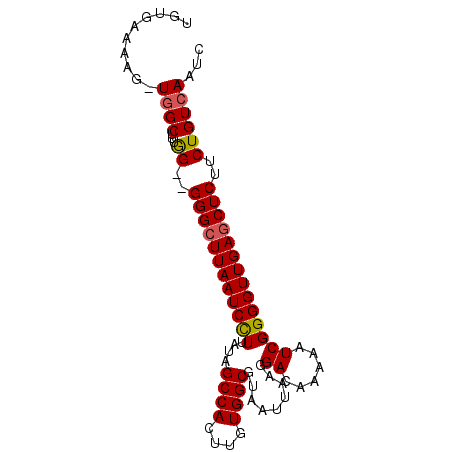
>3L_DroMel_CAF1 5385302 91 + 23771897 UGUGAAAAGCUUGCUUUGGGGGGGCUUAAUCCUUAUAGCCACUUGUGGCGUAAUUAAGGACAAAAAUCGGGGUUGAGCUCUUCUGUCAAUC ..(((((((....))))((((((((((((((((.((.((((....))))((........))....)).)))))))))))))))).)))... ( -30.40) >DroSec_CAF1 34450 88 + 1 UGUGAAAAG-UGGCUUUGG--GGGCUUAAUCCUUAUAGCCACUUGUGGCGUAAUUAAGGACGAAAAUCGGGGUUGAACUCUUCUGUCAAUC .((.(.(((-(((((.(((--(((.....)))))).)))))))).).)).........((((((....(((......)))))).))).... ( -25.10) >DroSim_CAF1 32692 88 + 1 UGUGAAAAG-UGGCUUUGG--GGGCUUAAUCCUUAUAGCCACUUGUGGCGCAAUUAAGGACAAAAAUCGGGGUUGAGCUCUGCUGUCAAUC ((((..(((-(((((.(((--(((.....)))))).))))))))....))))......((((.....(((((.....))))).)))).... ( -29.70) >DroEre_CAF1 34411 87 + 1 UGUGAAAAG-UGGCU-UAG--GGGCUUAAUCUUUAUAGCCACUUGUGGCGUAAUUAAGGACAAAAAUCGGGGUUGAGCUCUUCUGUCAAUC .........-((((.-.((--(((((((((((..((.((((....))))((........))....))..)))))))))))))..))))... ( -27.80) >DroYak_CAF1 33120 88 + 1 UGUGAAAAU-UGGCUCUGG--GGGCUUAAUCUUUAUAGCCACUUGUGGCGUAAUUAGGGACAAAAAUCGGGGUUGAGCUCCUCUGUCAAUC .......((-((((...((--(((((((((((..((.((((....))))((........))....))..)))))))))))))..)))))). ( -30.50) >consensus UGUGAAAAG_UGGCUUUGG__GGGCUUAAUCCUUAUAGCCACUUGUGGCGUAAUUAAGGACAAAAAUCGGGGUUGAGCUCUUCUGUCAAUC ..........((((...((..((((((((((((....((((....)))).........((......))))))))))))))..))))))... (-21.28 = -21.28 + -0.00)


| Location | 5,385,331 – 5,385,432 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 93.39 |
| Mean single sequence MFE | -24.02 |
| Consensus MFE | -21.58 |
| Energy contribution | -21.02 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.593480 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
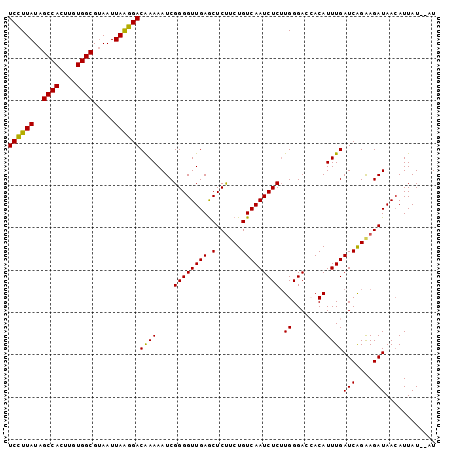
>3L_DroMel_CAF1 5385331 101 + 23771897 UCCUUAUAGCCACUUGUGGCGUAAUUAAGGACAAAAAUCGGGGUUGAGCUCUUCUGUCAAUCUCUUGGGACCACAUUUGAUCAGAAGAUAAAAUUAUAUAU ((((((..((((....)))).....))))))..........((((....(((((((((((.....((....))...)))).)))))))...))))...... ( -23.70) >DroSec_CAF1 34476 99 + 1 UCCUUAUAGCCACUUGUGGCGUAAUUAAGGACGAAAAUCGGGGUUGAACUCUUCUGUCAAUCUCUUGGGACCACAUUUGAUCAGAAGAUAACAUUAU--AU ((((((..((((....)))).....))))))((.....))..((((...(((((((((((.....((....))...)))).))))))))))).....--.. ( -25.60) >DroSim_CAF1 32718 99 + 1 UCCUUAUAGCCACUUGUGGCGCAAUUAAGGACAAAAAUCGGGGUUGAGCUCUGCUGUCAAUCUCUUGGGAGCACAUUUGAUCAGAAGAUAACAUUAU--AU ((((((..((((....)))).....)))))).....(((..(((..((...((((.((((....)))).))))..))..)))....)))........--.. ( -25.20) >DroEre_CAF1 34436 99 + 1 UCUUUAUAGCCACUUGUGGCGUAAUUAAGGACAAAAAUCGGGGUUGAGCUCUUCUGUCAAUCUCUUGGGACAACAUUUGAUCAGGAGAUAACAUUAU--AU ........((((....))))(((((....((((......((((.....))))..)))).(((((((((..((.....)).)))))))))...)))))--.. ( -21.90) >DroYak_CAF1 33146 99 + 1 UCUUUAUAGCCACUUGUGGCGUAAUUAGGGACAAAAAUCGGGGUUGAGCUCCUCUGUCAAUCUCGUGGGAGAACAUUUGAUCGGAAGAUAACAUUAU--AU ((((((..((((....)))).....)))))).....(((((((.....))))((((((((((((....))))....)))).)))).)))........--.. ( -23.70) >consensus UCCUUAUAGCCACUUGUGGCGUAAUUAAGGACAAAAAUCGGGGUUGAGCUCUUCUGUCAAUCUCUUGGGACCACAUUUGAUCAGAAGAUAACAUUAU__AU ((((((..((((....)))).....))))))((((....((((((((.(......))))))))).((......))))))(((....)))............ (-21.58 = -21.02 + -0.56)


| Location | 5,385,432 – 5,385,551 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 90.76 |
| Mean single sequence MFE | -32.69 |
| Consensus MFE | -23.64 |
| Energy contribution | -25.88 |
| Covariance contribution | 2.24 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.621048 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
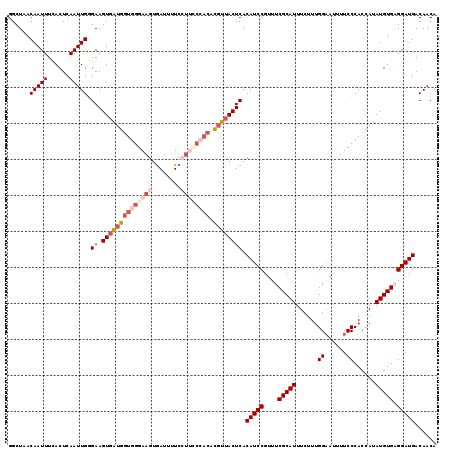
>3L_DroMel_CAF1 5385432 119 + 23771897 GGCUAACAAUUUCACUCAAUUGGGAAGUGAUGGUUGGAAGUGAUUUUCCUUCCCACACGUUACUCACAUCCGUUUCGCAUUUCUUUGGAAUUUUCCCACCAUAUGUGAGGAUGACAACA ......((.((((((.....((((((((((((((.(((((.(.....)))))).)).)))))).....((((.............))))...))))))......)))))).))...... ( -30.62) >DroSec_CAF1 34575 119 + 1 GGCUAACAAUUUCACUCAAUUGGGAAGUUAUGGUGGGAAGUGAUUUUCCUUCCCACACGUUACUCACAUCCGUUUCGCAUUUCUUUGGAAUUUUCCCACCAUAUGUGAGGAUGACAACA (..((((((((((.(......).))))))...((((((((.(.....)))))))))..))))..).(((((...((((((......(((....)))......)))))))))))...... ( -34.30) >DroSim_CAF1 32817 119 + 1 GGCUAACAAUUUCACUCAAUUGGGAAGUGCUGGUGGGAAGUGAUUUUCCUUUCCACACGGUACUCACAUCCGUUUCGCAUUUCUUCGGAAUUUUCCCACCAUAUGUGAGGAUGACAACA ......((.((((((.....((((((((((((((((((((.(.....))))))))).)))))).....((((.............))))...))))))......)))))).))...... ( -37.52) >DroEre_CAF1 34535 119 + 1 GGCUAACAAUUUUACUCAAUUGCGAAGUGAUGGUGGGAAGUGAUUUUCAUCGCCACACGUUACUCACAUCCGUUUCGCAUUUCUUUGGAAUUUUCCCACCAUAUGUGCGGAUGACAACA (..((((........((......)).(((.((((((((((....)))).)))))))))))))..).(((((((...((((......(((....)))......)))))))))))...... ( -30.00) >DroYak_CAF1 33245 118 + 1 GGCUAACAAUUUUACUCAAUUGCGAAGUGAUGGUGGGAAGUAAUGGUGGGAA-GUGAUUUUACUCACAUCCGUUUCGCAUUUCUUUGGCAUUUUCCCACCAUAUGUGUGGAUGACAACA ...............(((.(..((.....(((((((((((........(((.-((((......)))).))).....((.........))..)))))))))))...))..).)))..... ( -31.00) >consensus GGCUAACAAUUUCACUCAAUUGGGAAGUGAUGGUGGGAAGUGAUUUUCCUUCCCACACGUUACUCACAUCCGUUUCGCAUUUCUUUGGAAUUUUCCCACCAUAUGUGAGGAUGACAACA ......(((((......))))).((.((((((((((((((........)))))))).)))))))).(((((....(((((......((......))......))))).)))))...... (-23.64 = -25.88 + 2.24)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:20:13 2006