
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,348,555 – 5,348,694 |
| Length | 139 |
| Max. P | 0.837984 |

| Location | 5,348,555 – 5,348,656 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 90.31 |
| Mean single sequence MFE | -14.59 |
| Consensus MFE | -11.57 |
| Energy contribution | -11.45 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.837984 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
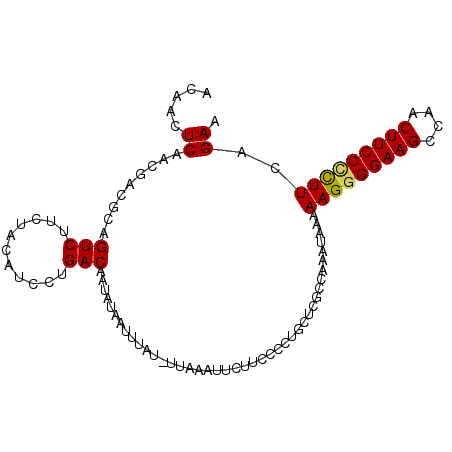
>3L_DroMel_CAF1 5348555 101 + 23771897 ACAACUCAACGACGCAGUCAUCAACAUCCUGACAAUAUAAUUUAU-UUAAAUUCUUCUCCUUCUCGCCAAAUAAAAAGAGGAAGCCAACUUCCCUUUCAGAA ..........(((...))).........((((........(((((-((....................)))))))(((.(((((....)))))))))))).. ( -10.75) >DroSec_CAF1 96377 102 + 1 ACAACUCAACGACGCAGUCUUCUACAUCCUGACAAUAUAUUUUUUUUUUCAAUCUUCCCCUGCUCACCAAAUAAAAAGGGGAAGCCAACUUCCUCUUCAGAA .....((.........(((...........)))....................((((((((...............))))))))...............)). ( -14.36) >DroSim_CAF1 98164 101 + 1 ACAACUCAACGACGCAGUCUUCUACAUCCUGACAAUAUAAUUUAU-UUAAAUUCUUCCCCUGCUCGCCAAAUAAAAAGGGGAAGCCAACUUCCUCUUCAGAA .....((..(((.((((((...........)))((((.....)))-).............)))))).........(((((((((....)))))))))..)). ( -16.70) >DroYak_CAF1 105084 101 + 1 ACAACUCAACAACGUAGUCUUCUACUUCCUGACAAUAUAAUUUAU-UUAAAUUCUACCCCUGCUCGCAAAAUAAAAAGGGGAAGCCAACUUCCCUUUCAGAA .............((((....))))...((((........(((((-((....................))))))).((((((((....)))))))))))).. ( -16.55) >consensus ACAACUCAACGACGCAGUCUUCUACAUCCUGACAAUAUAAUUUAU_UUAAAUUCUUCCCCUGCUCGCCAAAUAAAAAGGGGAAGCCAACUUCCCCUUCAGAA .....((.........(((...........)))..........................................(((((((((....)))))))))..)). (-11.57 = -11.45 + -0.12)


| Location | 5,348,594 – 5,348,694 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 81.49 |
| Mean single sequence MFE | -30.00 |
| Consensus MFE | -22.22 |
| Energy contribution | -22.14 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.606910 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
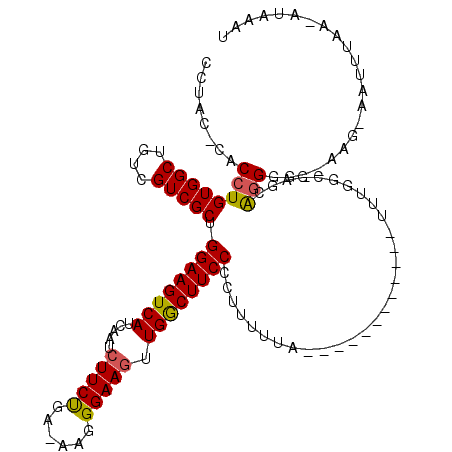
>3L_DroMel_CAF1 5348594 100 - 23771897 CCUAC--ACCUGUGGCUGUCGUCGCUGGAAGUCAUCAAUCUUCUGA-AAGGGAAGUUGGCUUCCUCUUUUUA------------UUUGGCGAGAAGGAG--AAG-AAUUUAA-AUAAAU .....--....(((((....))))).((((((((.....(((((..-...))))).))))))))(((((((.------------(((.....))).)))--)))-)......-...... ( -24.50) >DroSec_CAF1 96416 103 - 1 CCUACACACCUGUGGCUGUCGUCGCUGGAAGUCAUCAAUCUUCUGA-AGAGGAAGUUGGCUUCCCCUUUUUA------------UUUGGUGAGCAGGGG--AAG-AUUGAAAAAAAAAA ........((.(((((....))))).))......((((((((((..-...(((((....)))))(((.((((------------(...))))).)))))--)))-)))))......... ( -30.90) >DroSim_CAF1 98203 102 - 1 CCUACACACCUGUGGCUGUCGUCGCUGGAAGUCAUCAAUCUUCUGA-AGAGGAAGUUGGCUUCCCCUUUUUA------------UUUGGCGAGCAGGGG--AAG-AAUUUAA-AUAAAU (((.(((....))).((((..(((((((((((((....(((((...-.)))))...))))))))........------------...))))))))))))--...-.......-...... ( -29.00) >DroYak_CAF1 105123 99 - 1 -CUAC--ACCUGUGGCUGUCGUCGCUGGAAGUCAUCAAUCUUCUGA-AAGGGAAGUUGGCUUCCCCUUUUUA------------UUUUGCGAGCAGGGG--UAG-AAUUUAA-AUAAAU -((((--.((((((((....)))((.(((((........)))))((-((((((((....))).)))))))..------------....))..))))).)--)))-.......-...... ( -29.30) >DroAna_CAF1 98402 118 - 1 CCCACGCACCUGUGGCUGUCGUCGCUGGAAGUCAUCAAUCUUCCCAAAAGGGAACUUGACUUCCCCUUUCUCCCCGGAGGUCCUUUUCUCAAGCGCGGGAGUUGAAAAUAAA-AUAAAU (((.(((....(((((....))))).((((((((......(((((....)))))..))))))))....((((....))))............))).))).............-...... ( -36.30) >consensus CCUAC_CACCUGUGGCUGUCGUCGCUGGAAGUCAUCAAUCUUCUGA_AAGGGAAGUUGGCUUCCCCUUUUUA____________UUUGGCGAGCAGGGG__AAG_AAUUUAA_AUAAAU ........((((((((....))))).((((((((.....(((((......))))).))))))))..............................)))...................... (-22.22 = -22.14 + -0.08)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:19:47 2006