
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,174,344 – 5,174,436 |
| Length | 92 |
| Max. P | 0.983550 |

| Location | 5,174,344 – 5,174,436 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 92.05 |
| Mean single sequence MFE | -17.96 |
| Consensus MFE | -14.64 |
| Energy contribution | -14.88 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.773326 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5174344 92 + 23771897 AUCGCUGGCGUAAAUUUACGUAAAUAUUUAUAAGCUUUAGCACAACUUGACUGUCUCGAGAAUUCUUAAAGCCAAUAAGCACUUGCUGUGAA .((((..(((((....)))))............(((((((.....(((((.....))))).....))))))).....(((....))))))). ( -17.80) >DroSec_CAF1 39824 92 + 1 AUCACUGGCGUAAAUUUACUUAAAUAUUUAUAAGCUAUAGCACAACUUGACUUUCUCGAGAAUUCUUAAAGCCAAUAAGCGCUUGCUGUGAA .((((((((((((((..........))))))..((....))....(((((.....)))))..........))))...(((....))))))). ( -16.70) >DroSim_CAF1 39739 92 + 1 AUCACUGGCGUAAAUUUACGUAAAUAUUUAUAAGCUUUAGCACAACUUGACUUUCUCGAGAAUUCUUAAAGCCAAUAAGCGCUUGCUGUGAA .((((.(((((..((((....))))..((((..(((((((.....(((((.....))))).....)))))))..)))))))))....)))). ( -19.00) >DroEre_CAF1 38732 90 + 1 AUCACUAGCACCAAU--GCGUAAAUAUUUAUAAGCUUUAGCACAACUUGACUUUCUCGAGAAUUCUUAAAGCCAAUAAGCACUUGCUGUGAG .((((.((((....(--((((((....))))..(((((((.....(((((.....))))).....)))))))......)))..)))))))). ( -19.20) >DroYak_CAF1 38211 90 + 1 AUCACUUGCACAAAU--ACGUAAAUUUUUAUUAGCUUUAGCACAACUUGACUUUCUCGAGAAUUCUUAAAGCCAAUAAGCACUUGCUGUGAA .((((..((((....--..)).....((((((.(((((((.....(((((.....))))).....))))))).)))))).....)).)))). ( -17.10) >consensus AUCACUGGCGUAAAUUUACGUAAAUAUUUAUAAGCUUUAGCACAACUUGACUUUCUCGAGAAUUCUUAAAGCCAAUAAGCACUUGCUGUGAA .((((((((........................((....))....(((((.....)))))..........))))...(((....))))))). (-14.64 = -14.88 + 0.24)



| Location | 5,174,344 – 5,174,436 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 92.05 |
| Mean single sequence MFE | -21.72 |
| Consensus MFE | -18.46 |
| Energy contribution | -18.58 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.95 |
| SVM RNA-class probability | 0.983550 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5174344 92 - 23771897 UUCACAGCAAGUGCUUAUUGGCUUUAAGAAUUCUCGAGACAGUCAAGUUGUGCUAAAGCUUAUAAAUAUUUACGUAAAUUUACGCCAGCGAU ......((.((..(...(((((((..((....))..)...))))))...)..))..................((((....))))...))... ( -14.70) >DroSec_CAF1 39824 92 - 1 UUCACAGCAAGCGCUUAUUGGCUUUAAGAAUUCUCGAGAAAGUCAAGUUGUGCUAUAGCUUAUAAAUAUUUAAGUAAAUUUACGCCAGUGAU .((((.((.(((((...((((((((..((....))...))))))))...)))))...(((((........)))))........))..)))). ( -24.90) >DroSim_CAF1 39739 92 - 1 UUCACAGCAAGCGCUUAUUGGCUUUAAGAAUUCUCGAGAAAGUCAAGUUGUGCUAAAGCUUAUAAAUAUUUACGUAAAUUUACGCCAGUGAU .(((((((.(((((...((((((((..((....))...))))))))...)))))...)))............((((....))))...)))). ( -23.70) >DroEre_CAF1 38732 90 - 1 CUCACAGCAAGUGCUUAUUGGCUUUAAGAAUUCUCGAGAAAGUCAAGUUGUGCUAAAGCUUAUAAAUAUUUACGC--AUUGGUGCUAGUGAU .((((((((((..(...((((((((..((....))...))))))))...)..))...((.((........)).))--.....)))).)))). ( -23.70) >DroYak_CAF1 38211 90 - 1 UUCACAGCAAGUGCUUAUUGGCUUUAAGAAUUCUCGAGAAAGUCAAGUUGUGCUAAAGCUAAUAAAAAUUUACGU--AUUUGUGCAAGUGAU .((((.((((((((...((((((((..((....))...))))))))((((.((....))))))..........))--))))))....)))). ( -21.60) >consensus UUCACAGCAAGUGCUUAUUGGCUUUAAGAAUUCUCGAGAAAGUCAAGUUGUGCUAAAGCUUAUAAAUAUUUACGUAAAUUUACGCCAGUGAU .(((((((.(((((...((((((((..((....))...))))))))...)))))...)))............((((....))))...)))). (-18.46 = -18.58 + 0.12)

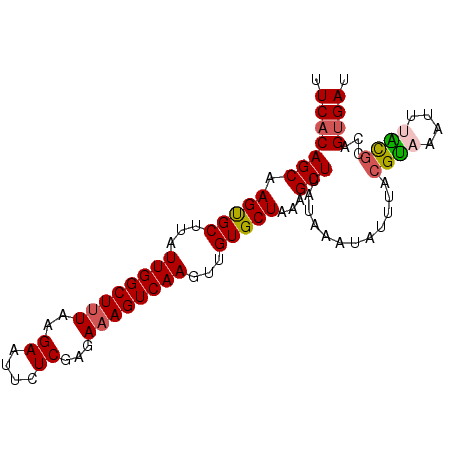

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:18:10 2006