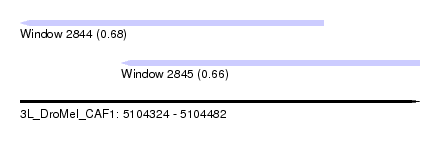

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,104,324 – 5,104,482 |
| Length | 158 |
| Max. P | 0.682272 |

| Location | 5,104,324 – 5,104,444 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -31.27 |
| Consensus MFE | -27.21 |
| Energy contribution | -27.90 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.682272 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
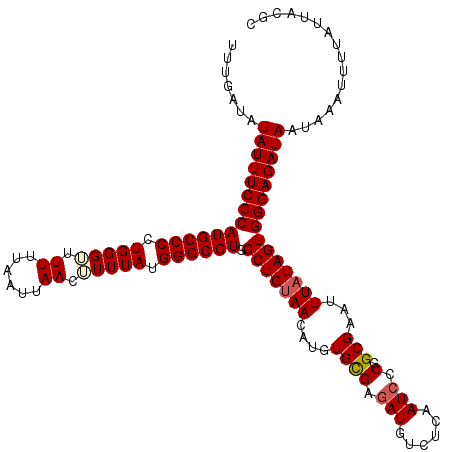
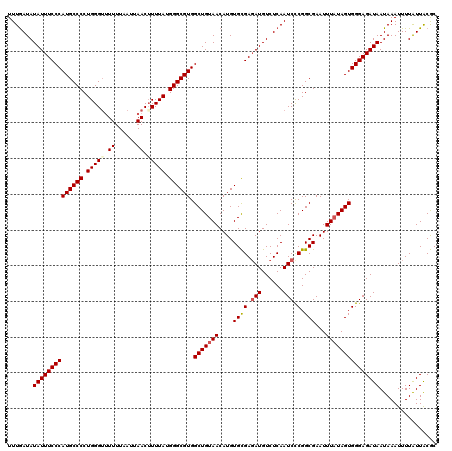
>3L_DroMel_CAF1 5104324 120 - 23771897 UUUGAUAUAUUUCCCAUGCCCCUGGGUUUUUUAAUUAACAUUUAUGGGCGUGGCUGUAACAUGUGCGAUAUGUCUCAAUCCCGGCGAAUUUUUAGUGGGAGAUAAUAAAUUUUAUUACGC ...((((((((..((((((((.(((((.((......)).))))).))))))))..(((.....)))))))))))....(((((.(.........))))))(((((......))))).... ( -28.10) >DroSec_CAF1 297269 120 - 1 UUUAAUAUAUUUCCCAUGCCCCUGGGUUUUUUAAUUAACUUUUAUGGGCGUGGCUGUAACAUGUGCGAGAUGUCUCAAUCCCGGCGAAUUUAUAGUGGGAGAUAAUAAAUUUUGUUACGC .............((((((((..(((((........)))))....))))))))..((((((.....(((....))).(((((.((.........)).)).))).........)))))).. ( -32.00) >DroSim_CAF1 318922 120 - 1 UUUGAUAUAUUUCCCAUGCCCCUGGGUUUUUUAAUUAACUUUUAUGGGCGUGGCUGUAACAUGUGCGAGAUGUCUCAAUCCCGGCGAAUUUAUAGUGGGAGAUAAUAAAUUUUAUUACGC .......((((((((((((((..(((((........)))))....)))))).(((((((....((((.(((......))).).)))...)))))))))))))))................ ( -31.60) >DroYak_CAF1 309600 120 - 1 UUUGAUAUAUUUCCCAUGCCCCUGGGACUUUUAAUUAACUUUUAUGGGCGUGGCUGUAACAUGUGUGAGAUGUCUCAAUCUCGGCGAAUUUAUAGUGGGAGAUAAUAAAUUUUAUUGCAC .......((((((((((((((.(((((.((......)).))))).)))))).(((((((....((((((((......))))).)))...)))))))))))))))................ ( -33.40) >consensus UUUGAUAUAUUUCCCAUGCCCCUGGGUUUUUUAAUUAACUUUUAUGGGCGUGGCUGUAACAUGUGCGAGAUGUCUCAAUCCCGGCGAAUUUAUAGUGGGAGAUAAUAAAUUUUAUUACGC .......((((((((((((((.(((((.((......)).))))).)))))).(((((((....((((.(((......))).).)))...)))))))))))))))................ (-27.21 = -27.90 + 0.69)

| Location | 5,104,364 – 5,104,482 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.15 |
| Mean single sequence MFE | -30.20 |
| Consensus MFE | -19.64 |
| Energy contribution | -20.33 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.660310 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
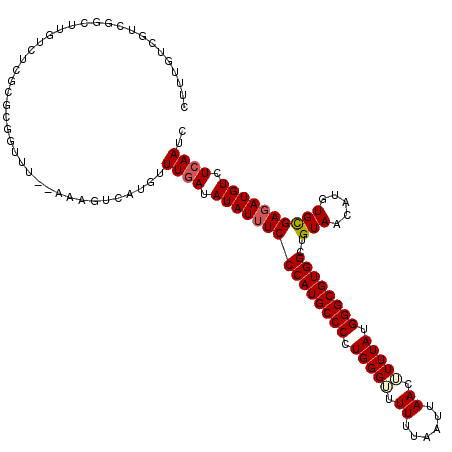
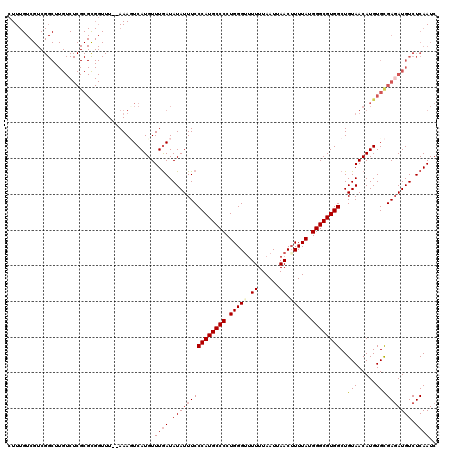
>3L_DroMel_CAF1 5104364 118 - 23771897 CUUUGUCGUCGGCUUGUCUCGCGCGGUUU--AAAUUCAUGUUUGAUAUAUUUCCCAUGCCCCUGGGUUUUUUAAUUAACAUUUAUGGGCGUGGCUGUAACAUGUGCGAUAUGUCUCAAUC ..........(((.....(((((((..((--((((....))))))..(((...((((((((.(((((.((......)).))))).))))))))..)))...)))))))...)))...... ( -28.80) >DroSec_CAF1 297309 93 - 1 CUU-------------------------U--AAAGUCAUGUUUAAUAUAUUUCCCAUGCCCCUGGGUUUUUUAAUUAACUUUUAUGGGCGUGGCUGUAACAUGUGCGAGAUGUCUCAAUC ...-------------------------.--...(((((((....((((....((((((((..(((((........)))))....)))))))).))))))))).))(((....))).... ( -23.80) >DroSim_CAF1 318962 118 - 1 CUUUGUCGUCGGCUUGUCUCGCGCGGUUU--AAAGUCAUGUUUGAUAUAUUUCCCAUGCCCCUGGGUUUUUUAAUUAACUUUUAUGGGCGUGGCUGUAACAUGUGCGAGAUGUCUCAAUC ..........(((..(((((((((((((.--...((((....)))).......((((((((..(((((........)))))....))))))))....))).)))))))))))))...... ( -36.90) >DroYak_CAF1 309640 120 - 1 CUUUGUCGUCGGCUUGUCUCGCGUGGUUUUAAAACUCAUGUUUGAUAUAUUUCCCAUGCCCCUGGGACUUUUAAUUAACUUUUAUGGGCGUGGCUGUAACAUGUGUGAGAUGUCUCAAUC ...(((...((((.((((..((((((((....))).)))))..)))).......(((((((.(((((.((......)).))))).)))))))))))..)))....((((....))))... ( -31.30) >consensus CUUUGUCGUCGGCUUGUCUCGCGCGGUUU__AAAGUCAUGUUUGAUAUAUUUCCCAUGCCCCUGGGUUUUUUAAUUAACUUUUAUGGGCGUGGCUGUAACAUGUGCGAGAUGUCUCAAUC .........................................((((.(((((((((((((((.(((((.((......)).))))).))))))))..(((.....)))))))))).)))).. (-19.64 = -20.33 + 0.69)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:17:38 2006