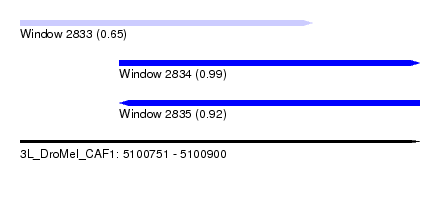
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,100,751 – 5,100,900 |
| Length | 149 |
| Max. P | 0.986303 |

| Location | 5,100,751 – 5,100,860 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.52 |
| Mean single sequence MFE | -25.52 |
| Consensus MFE | -17.42 |
| Energy contribution | -17.93 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.648947 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5100751 109 + 23771897 UACACCAUG-CCACUAUAU--AUUUCGUACACUUCUUGAUGUCGCCCAAGUGGGCGUGGCCAGCGCAAAAGCAA------AAGAUCCCAAGUGCUAAAUCAACAAA--ACAAAAGAACAA ........(-((((.....--.....(.((((.....).))))((((....))))))))).(((((........------..........)))))...........--............ ( -20.87) >DroSec_CAF1 293827 117 + 1 UACAUCUUG-CCACUAUAUACAUUUGGUACUCCUCUUCAUGUCGCCCAAGUGGGCGUGGCCAGCGCCAAAGCAAAAGCACAAGAUCCCAAGUGCUAAAUCAACAAA--ACAAAAGAACAA ....((((.-...........((((((((((.....((.((((((((....)))))((((....))))..((....))))).)).....)))))))))).......--....)))).... ( -28.00) >DroSim_CAF1 315420 117 + 1 UACAUCUUG-CCACUAUAUAUAUUUGGUACACCUCUUCAUGUCGCCCAAGUGGGCGUGGCCAGCGCCAAAGCAAAAGCACAAGAUCCCAAGUGCUAAAUCAACAAA--ACAAAAGAACAA ....(((((-((((...........((....))..........((((....)))))))))..((......))...(((((..........)))))...........--....)))).... ( -26.60) >DroYak_CAF1 306121 114 + 1 UACACUCUGCCCACUAUAUAUAUUUGGUAAGCCUCUUCAUGUCGCCCAAGUGGGCGUGGCUGGCGUAAUGGCAA------AAGAUUCCAAGUGCUAAAUCAACAAAGAGCAAAAGAACAA ....(((((((.((((........)))).((((.........(((((....))))).)))))))....(((((.------..(....)...))))).........))))........... ( -26.60) >consensus UACACCUUG_CCACUAUAUAUAUUUGGUACACCUCUUCAUGUCGCCCAAGUGGGCGUGGCCAGCGCAAAAGCAA______AAGAUCCCAAGUGCUAAAUCAACAAA__ACAAAAGAACAA .....................(((((((((............(((((....)))))......((......))..................)))))))))..................... (-17.42 = -17.93 + 0.50)



| Location | 5,100,788 – 5,100,900 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.38 |
| Mean single sequence MFE | -36.37 |
| Consensus MFE | -30.68 |
| Energy contribution | -30.12 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.34 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.04 |
| SVM RNA-class probability | 0.986303 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5100788 112 + 23771897 GUCGCCCAAGUGGGCGUGGCCAGCGCAAAAGCAA------AAGAUCCCAAGUGCUAAAUCAACAAA--ACAAAAGAACAACGACCGAUACAAAUUUGUUCGCUGCUGCGCUGCCUUUAAU ..(((((....))))).(((.((((((..(((..------..........((.........))...--......((((((..............)))))))))..)))))))))...... ( -33.04) >DroSec_CAF1 293866 118 + 1 GUCGCCCAAGUGGGCGUGGCCAGCGCCAAAGCAAAAGCACAAGAUCCCAAGUGCUAAAUCAACAAA--ACAAAAGAACAAAGACCGAUACAAAUUUGUUCGCUGCUGCGCUGCCUUUAAU ..(((((....))))).(((.(((((...(((...(((((..........)))))...........--......(((((((............))))))))))...))))))))...... ( -39.10) >DroSim_CAF1 315459 118 + 1 GUCGCCCAAGUGGGCGUGGCCAGCGCCAAAGCAAAAGCACAAGAUCCCAAGUGCUAAAUCAACAAA--ACAAAAGAACAACGACCGAUACAAAUUUGUUCGCUGCUGCGCUGCCUUUAAU ..(((((....))))).(((.(((((...(((...(((((..........)))))...........--......((((((..............)))))))))...))))))))...... ( -37.14) >DroYak_CAF1 306161 114 + 1 GUCGCCCAAGUGGGCGUGGCUGGCGUAAUGGCAA------AAGAUUCCAAGUGCUAAAUCAACAAAGAGCAAAAGAACAACGAGCGAUACAAAUUUGUUCGCUGCUGCGCUGCCUUUAAU ..(((((....))))).(((.((((((..(((..------....(((....((((............))))...)))....((((((.......)))))))))..)))))))))...... ( -36.20) >consensus GUCGCCCAAGUGGGCGUGGCCAGCGCAAAAGCAA______AAGAUCCCAAGUGCUAAAUCAACAAA__ACAAAAGAACAACGACCGAUACAAAUUUGUUCGCUGCUGCGCUGCCUUUAAU ..(((((....))))).(((.(((((...(((..................((.........))...........((((((..............)))))))))...))))))))...... (-30.68 = -30.12 + -0.56)



| Location | 5,100,788 – 5,100,900 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.38 |
| Mean single sequence MFE | -37.42 |
| Consensus MFE | -31.84 |
| Energy contribution | -33.15 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.922433 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5100788 112 - 23771897 AUUAAAGGCAGCGCAGCAGCGAACAAAUUUGUAUCGGUCGUUGUUCUUUUGU--UUUGUUGAUUUAGCACUUGGGAUCUU------UUGCUUUUGCGCUGGCCACGCCCACUUGGGCGAC ......((((((((((.(((((((((..(((...)))...))))))......--......(((((........)))))..------..))).))))))).))).(((((....))))).. ( -37.80) >DroSec_CAF1 293866 118 - 1 AUUAAAGGCAGCGCAGCAGCGAACAAAUUUGUAUCGGUCUUUGUUCUUUUGU--UUUGUUGAUUUAGCACUUGGGAUCUUGUGCUUUUGCUUUGGCGCUGGCCACGCCCACUUGGGCGAC ......((((((((...((((((((((............)))))))......--...........(((((..(....)..)))))...)))...))))).))).(((((....))))).. ( -39.50) >DroSim_CAF1 315459 118 - 1 AUUAAAGGCAGCGCAGCAGCGAACAAAUUUGUAUCGGUCGUUGUUCUUUUGU--UUUGUUGAUUUAGCACUUGGGAUCUUGUGCUUUUGCUUUGGCGCUGGCCACGCCCACUUGGGCGAC ......((((((((...(((((((((..(((...)))...))))))......--...........(((((..(....)..)))))...)))...))))).))).(((((....))))).. ( -39.30) >DroYak_CAF1 306161 114 - 1 AUUAAAGGCAGCGCAGCAGCGAACAAAUUUGUAUCGCUCGUUGUUCUUUUGCUCUUUGUUGAUUUAGCACUUGGAAUCUU------UUGCCAUUACGCCAGCCACGCCCACUUGGGCGAC ......(((.(((((((((((((((....))).))))).))))((((..((((............))))...))))....------.........)))..))).(((((....))))).. ( -33.10) >consensus AUUAAAGGCAGCGCAGCAGCGAACAAAUUUGUAUCGGUCGUUGUUCUUUUGU__UUUGUUGAUUUAGCACUUGGGAUCUU______UUGCUUUGGCGCUGGCCACGCCCACUUGGGCGAC ......((((((((((.(((((((((..(((...)))...))))))..............(((((........)))))..........))).))))))).))).(((((....))))).. (-31.84 = -33.15 + 1.31)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:17:28 2006