
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 5,083,690 – 5,083,841 |
| Length | 151 |
| Max. P | 0.813716 |

| Location | 5,083,690 – 5,083,801 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.13 |
| Mean single sequence MFE | -30.24 |
| Consensus MFE | -22.36 |
| Energy contribution | -22.68 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.813716 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5083690 111 - 23771897 AAGA-CAGCGUA-------GAGAGAAGAGAGAAAUGCAGAUUGCAUGUCCUGGCAACAGGAUGCACCCGUCCCAGUCAUGCAUUAAUAACAACCCCAU-CUCCAGCAAACAAUCCCAGCC ....-..((...-------((((.........(((((((((((...((((((....))))))((....))..))))).)))))).............)-)))..)).............. ( -25.05) >DroSec_CAF1 289909 109 - 1 AAGA-CAGCGAG---------GAGAAGAGGGAAAUGCAGAUUGCAUGUCCUGGCAACAGGAUGCACCCGCCCCAGUCAUGCAUUAACAACAACGCCAU-CUCCAGCAAACAAUCCCAGCC ....-..((..(---------((((....((.(((((((((((...((((((....))))))((....))..))))).))))))..........)).)-)))).)).............. ( -31.30) >DroSim_CAF1 311210 109 - 1 AAGA-CAGCGAA---------GAGAAGAGGGAAAUGCAGAUUGCAUGUCCUGGCAACAGGAUGCACCCGCCCCAGUCAUGCAUUAACAACAACGCCAU-CUCCAGCAAACAAUCCCAGCC ....-.......---------.......(((((((((((((((...((((((....))))))((....))..))))).)))))).........((...-.....))......)))).... ( -29.80) >DroYak_CAF1 301464 120 - 1 AAAAGCAGCGAGAGGCUGACUGAAAUGAGAGAAAUGCAGAUUGUAUGUCCUGGCAACAGGAUGCAUUCGCCCCAGUCAUGCAUUAACAACAACCCCAUACCCCAGCAAACAAUCGCAGCC ....((.((((...((((.((......))...(((((((((((...((((((....))))))((....))..))))).))))))..................))))......)))).)). ( -34.80) >consensus AAGA_CAGCGAA_________GAGAAGAGAGAAAUGCAGAUUGCAUGUCCUGGCAACAGGAUGCACCCGCCCCAGUCAUGCAUUAACAACAACCCCAU_CUCCAGCAAACAAUCCCAGCC ............................(((((((((((((((...((((((....))))))((....))..))))).))))))....................(....)..)))).... (-22.36 = -22.68 + 0.31)



| Location | 5,083,729 – 5,083,841 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.63 |
| Mean single sequence MFE | -33.72 |
| Consensus MFE | -27.48 |
| Energy contribution | -27.10 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.81 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 5083729 112 - 23771897 UUUGGCAAAAGUUGCCUACGAUCUUUUCGGACUACAAACGAAGA-CAGCGUA-------GAGAGAAGAGAGAAAUGCAGAUUGCAUGUCCUGGCAACAGGAUGCACCCGUCCCAGUCAUG ..((((....((((((.....(((((((...((((.........-....)))-------)...)))))))((.((((.....)))).))..)))))).(((((....)))))..)))).. ( -32.92) >DroSec_CAF1 289948 110 - 1 UUUGGCAAAAGUUGCCUACGAUCUUUUCGGACUACAAACGAAGA-CAGCGAG---------GAGAAGAGGGAAAUGCAGAUUGCAUGUCCUGGCAACAGGAUGCACCCGCCCCAGUCAUG ...(((........(((.((...((((((.........))))))-...))))---------)......(((..((((.....))))((((((....))))))...))))))......... ( -34.40) >DroSim_CAF1 311249 110 - 1 UUUGGCAAAAGUUGCCUACGAUCUUUUCGGACUACAAACGAAGA-CAGCGAA---------GAGAAGAGGGAAAUGCAGAUUGCAUGUCCUGGCAACAGGAUGCACCCGCCCCAGUCAUG ...(((....((((((.....((((((((...............-...))))---------))))...((((.((((.....)))).)))))))))).((......)))))......... ( -32.97) >DroYak_CAF1 301504 120 - 1 UUUGGCAAAAGUUGCCUACGAUCUCUUCGGACUACAAGCGAAAAGCAGCGAGAGGCUGACUGAAAUGAGAGAAAUGCAGAUUGUAUGUCCUGGCAACAGGAUGCAUUCGCCCCAGUCAUG ...((((.....))))..(((.....)))((((....(((((.((((((.....)))).))............((((.....))))((((((....))))))...)))))...))))... ( -34.60) >consensus UUUGGCAAAAGUUGCCUACGAUCUUUUCGGACUACAAACGAAGA_CAGCGAA_________GAGAAGAGAGAAAUGCAGAUUGCAUGUCCUGGCAACAGGAUGCACCCGCCCCAGUCAUG ...((((.....))))..(((.....)))((((.....((........))..................(((..((((.....))))((((((....))))))...))).....))))... (-27.48 = -27.10 + -0.38)

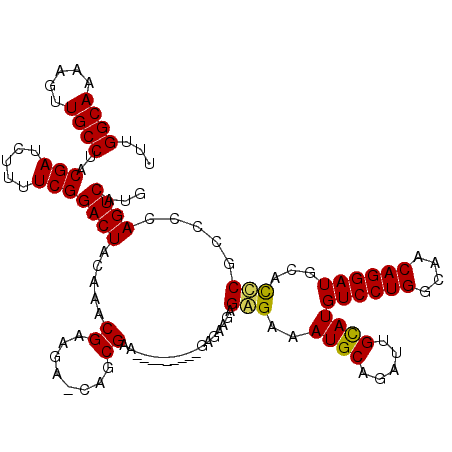
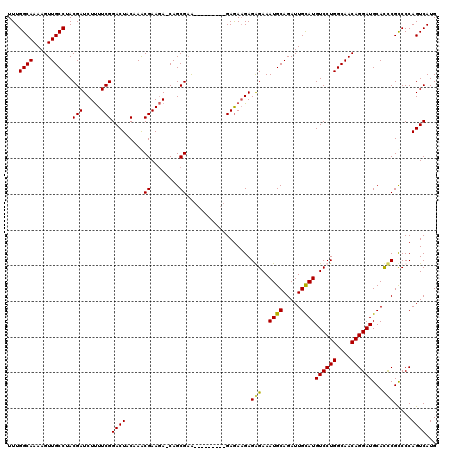
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:17:16 2006